Exploring zebrafish genes and transcripts, demo
Demo: The gene tab
If you click on any one of the transcripts in the Region in detail image, a pop-up menu will appear, allowing you to jump directly to that gene or transcript.
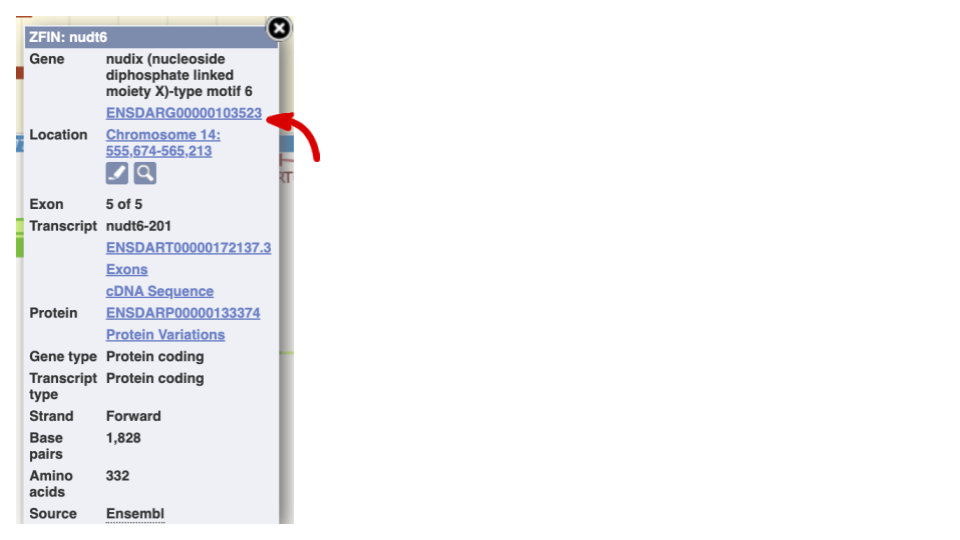
Another way to go to a gene of interest is to search directly for it.
We’re going to look at the zebrafish espn gene. This gene encodes a multifunctional actin-bundling protein with a major role in mediating sensory transduction in various mechanosensory and chemosensory cells. Mutations in this gene are associated with deafness in humans.
From ensembl.org, type espn into the search bar and click the Go button. You will get a list of hits with the human gene at the top.
Where you search for something without specifying the species, or where the ID is not restricted to a single species, the most popular species will appear first, in this case, human, mouse and zebrafish appear first. You can restrict your query to species or features of interest using the options on the left.
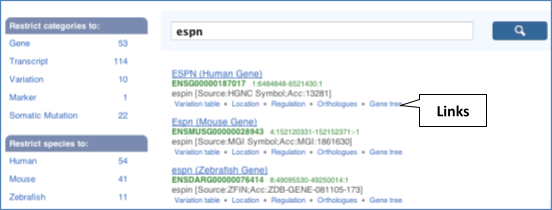
Click on the gene name of the zebrafish gene. The Gene tab should open:
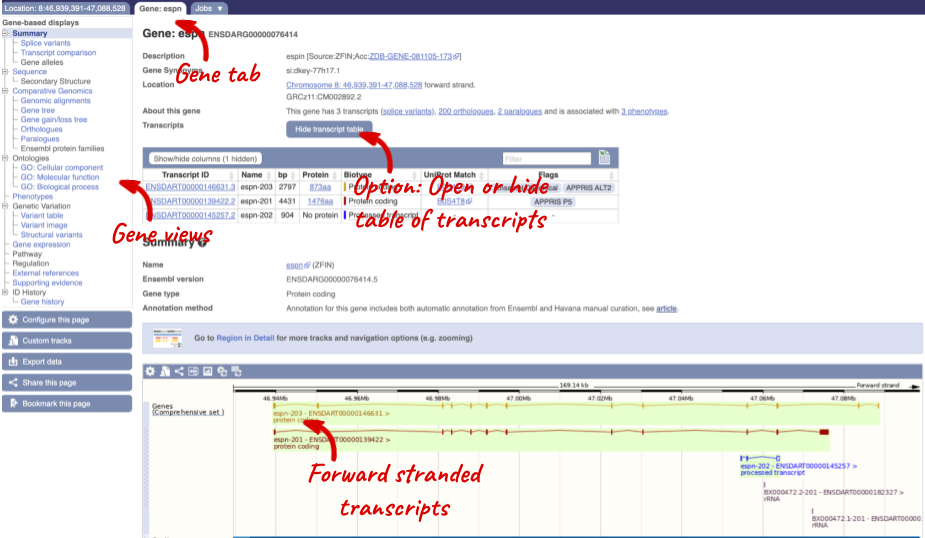
Let’s walk through some of the links in the left hand navigation column. How can we view the genomic sequence? Click Sequence at the left of the page.
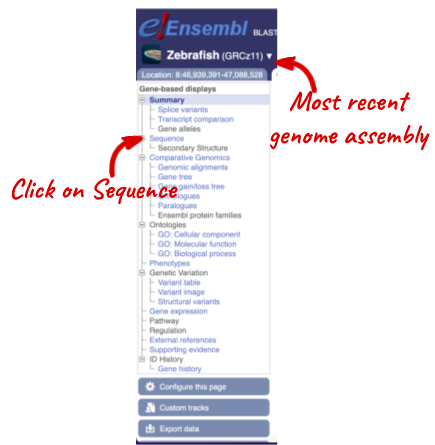

The sequence is shown in FASTA format. Take a look at the FASTA header:
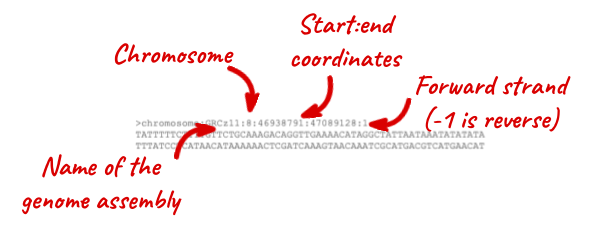
Exons are highlighted within the genomic sequence. Variants can be added with the Configure this page link found at the left. Click on it now.
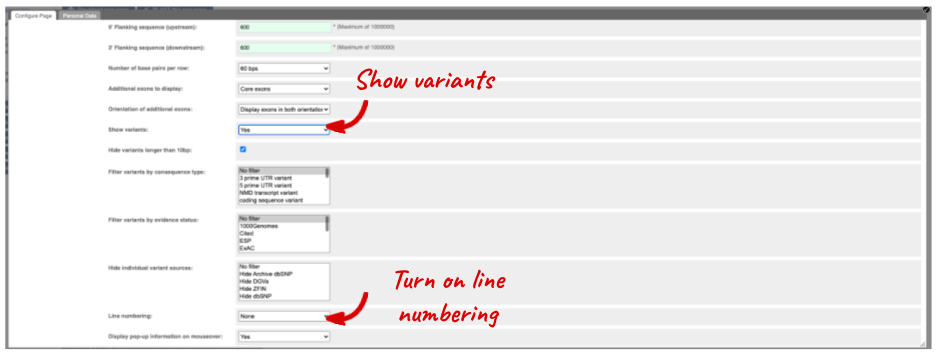
Once you have selected changes (in this example, Show variants and Line numbering) click at the top right.
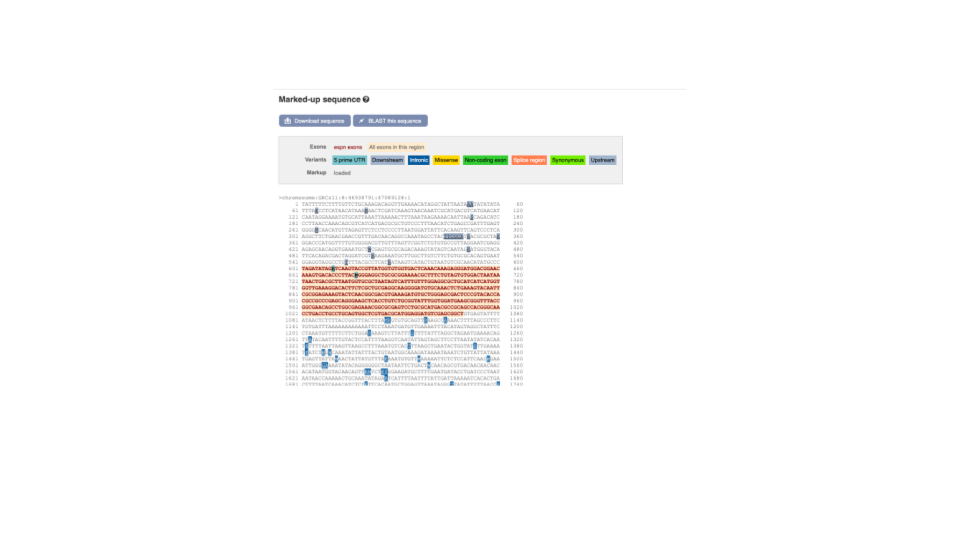
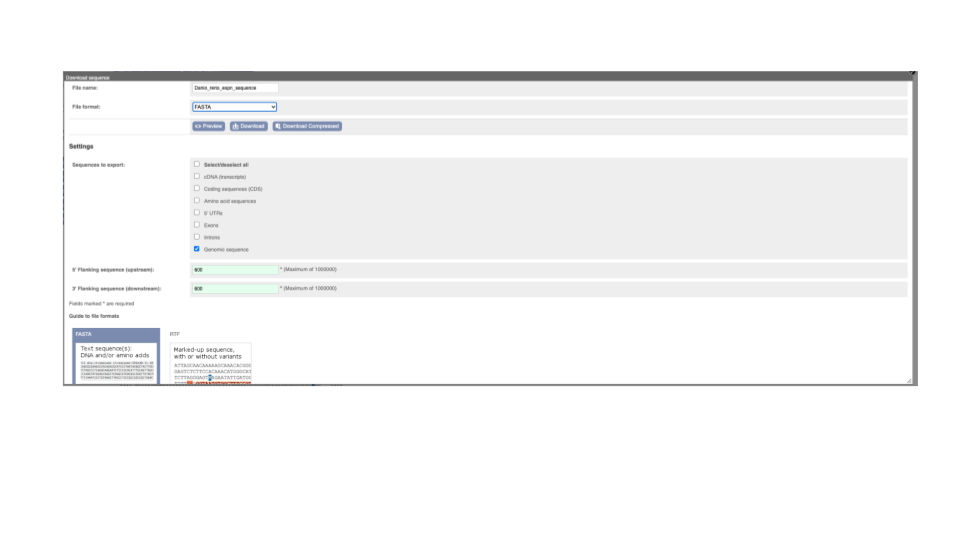
You can download this sequence by clicking in the Download sequence button above the sequence:

This will open a dialogue box that allows you to pick between plain FASTA sequence, or sequence in RTF, which includes all the coloured annotations and can be opened in a word processor. This button is available for all sequence views.
Can our gene be found in other databases? Go up the left-hand menu to External references:
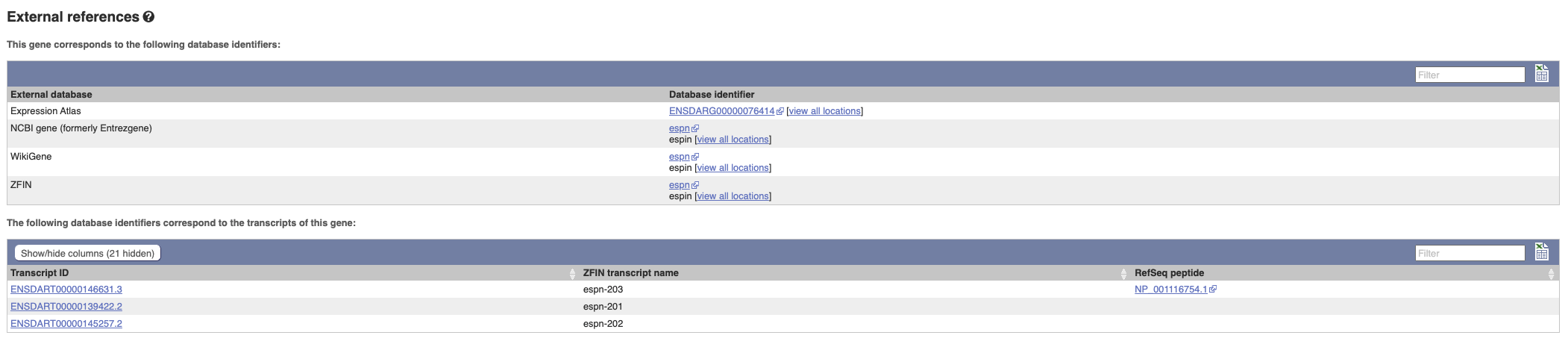
This contains links to the gene in other projects, such as NCBI Gene, and papers where this sequence is published.
To find out what the protein does, click on GO:biological process to see GO terms from the Gene Ontology consortium.
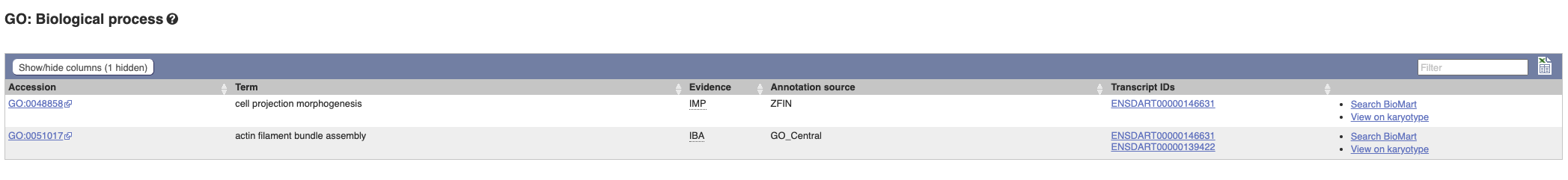
Hover over the three-letter Evidence codes to find out what they mean.
Demo: The transcript tab
Let’s now explore one splice isoform. Click on Show transcript table at the top.

Click on the ID for espn-203, ENSDART00000146631.3.
You are now in the Transcript tab for espn-203. The left hand navigation column provides several options for the transcript espn-203.
For detailed information on the support for this transcript, click on Supporting evidence
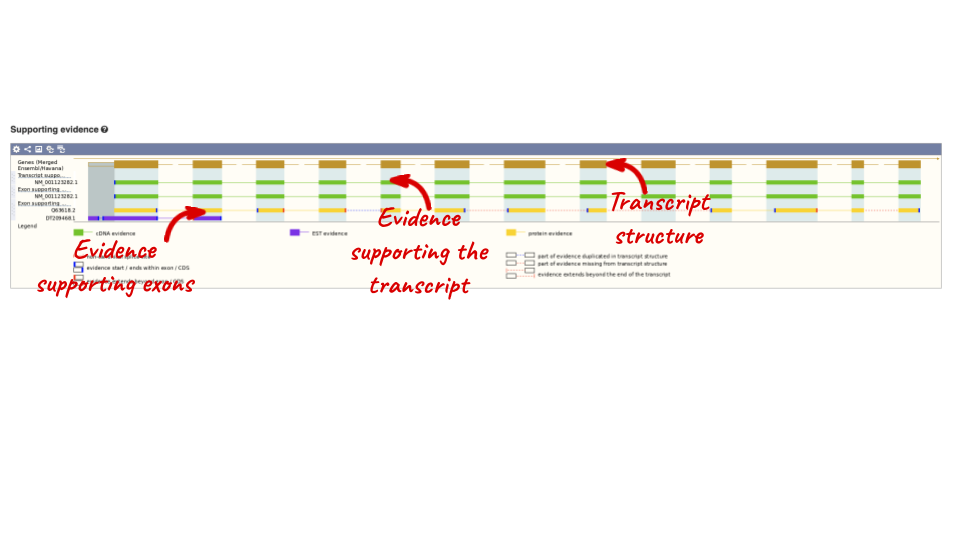
Click on the identifiers of the evidence to get a pop-up. This links out to the original records of these data in, for example, RefSeq, Uniprot or ENA.
Click on the _Exons _link.
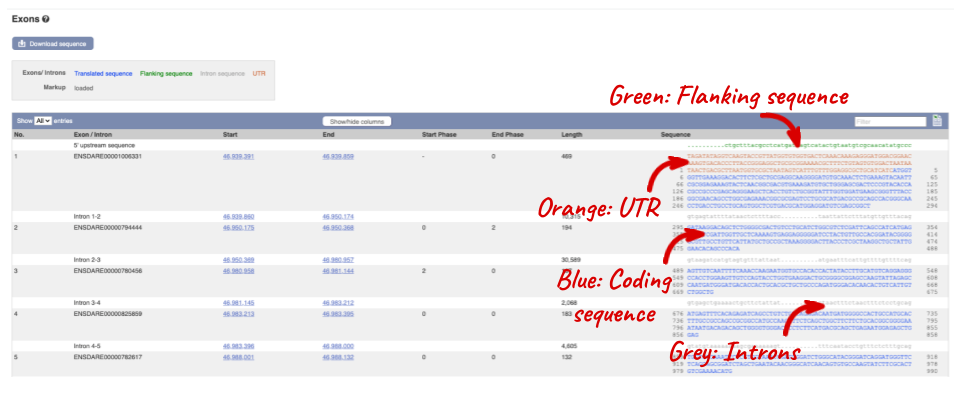
You may want to change the display (for example, to show more flanking sequence, or to show full introns). In order to do so click on Configure this page and change the display options accordingly.
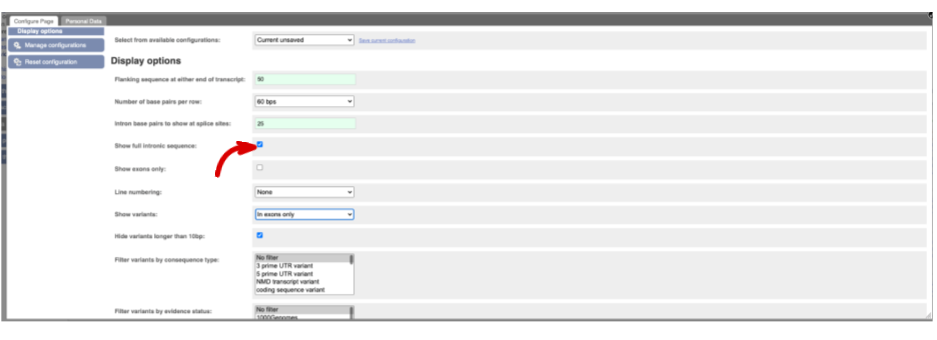
Now click on the_ cDNA_ link to see the spliced transcript sequence.
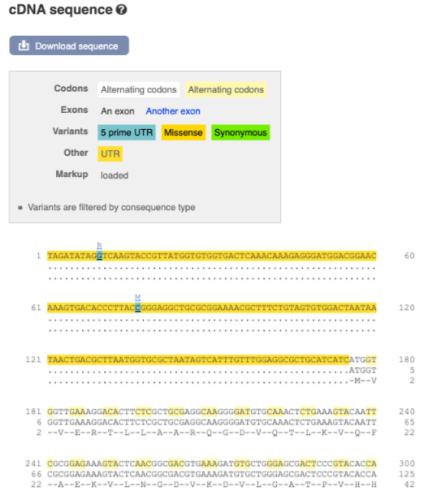
UnTranslated Regions (UTRs) are highlighted in dark yellow, codons are highlighted in light yellow, and exon sequence is shown in black or blue letters to show exon divides. Sequence variants are represented by highlighted nucleotides and clickable IUPAC codes are above the sequence.
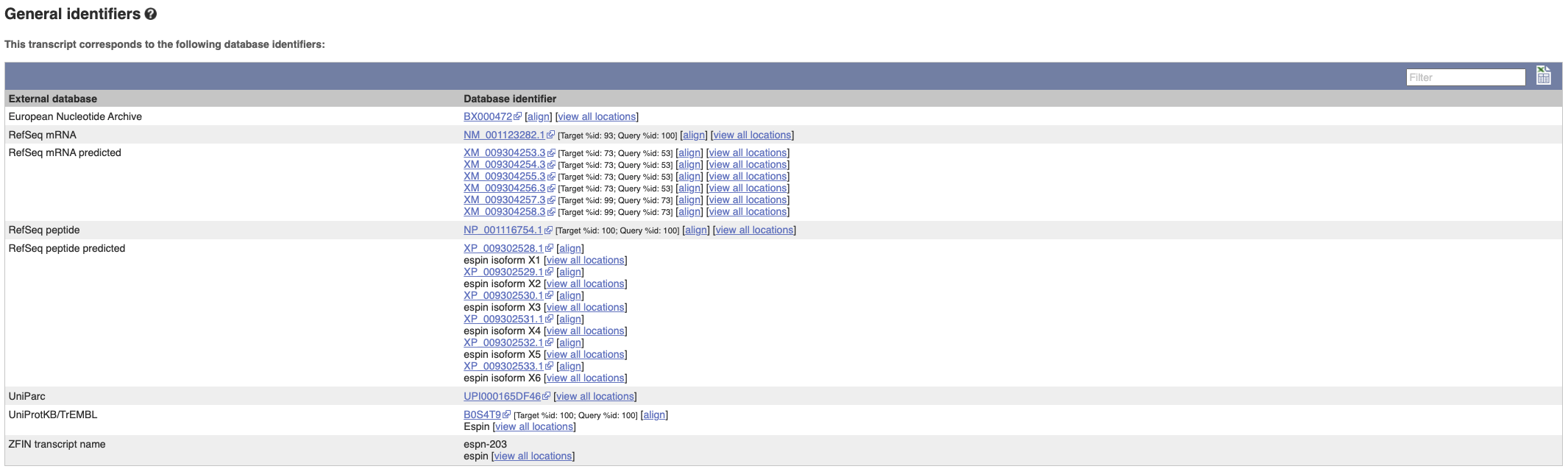
Next, follow the General identifiers link at the left.
This page shows information from other databases such as RefSeq, UniProtKB and others, that match to the Ensembl transcript and protein.
Now click on Protein summary to view domains from Pfam, PROSITE, Superfamily, InterPro, and more.
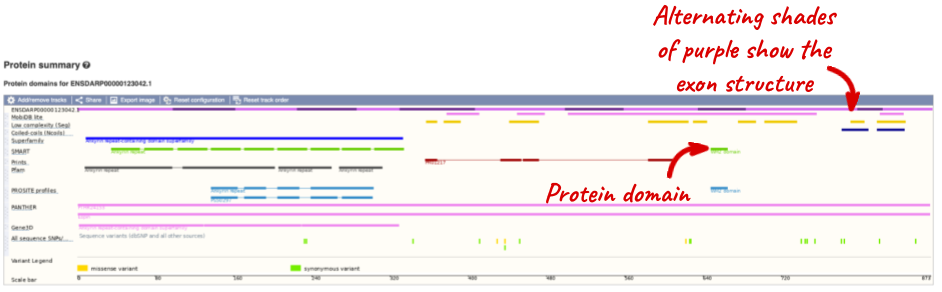
Clicking on Domains & features shows a table of this information.






