Annotating genetic variants with the VEP, Demo
We have identified three variants on sheep chromosome one, G->A at 193980041, G->T at 194005540 and a C deletion at 194033920.
We will use the Ensembl VEP to determine:
- If the variants have been annotated in Ensembl already
- If genes are affected by the variants
Go to the front page of Ensembl and click on Variant Effect Predictor in the Tools section or click on VEP in the top header.

This page contains information about the VEP, including a link for downloading the script version of the tool. Click on the Launch VEP button to open the input form.
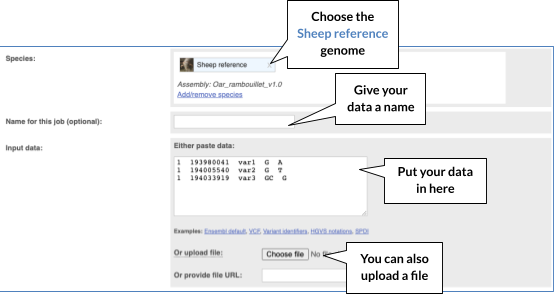
The data is in VCF format:
Chromosome Position Name Reference Alternative
Put the following into the Input data box:
1 193980041 var1 G A
1 194005540 var2 G T
1 194033919 var3 GC G
The VEP will detect automatically that the data is in VCF format.
There are further options that you can choose for your output. These are categorised as Identifiers and frequency data, Extra options and Filtering options. Let’s open all menus and take a look.
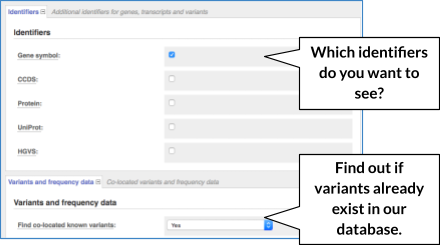
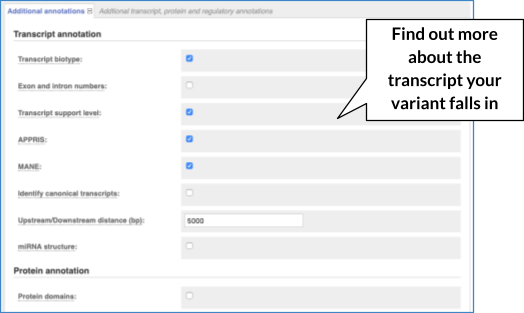
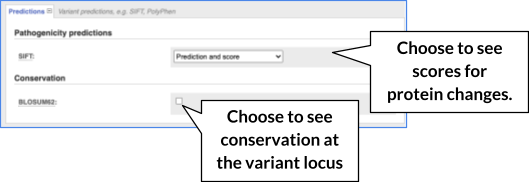
Hover over the options to see definitions.
When you have selected everything you need, scroll right to the bottom and click Run.

The display will show you the status of your job. It will say Queued, then automatically switch to Done when the job is done, you do not need to refresh the page. You can save, edit, share or delete your job at this time. If you have submitted multiple jobs, they will all appear here.
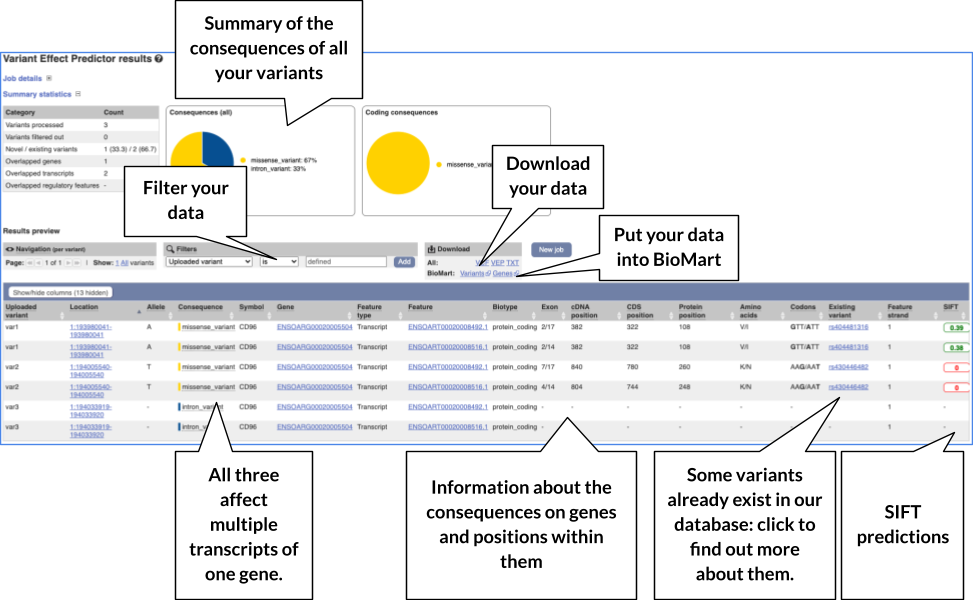
Click on View Results once your job is done.
In your results you will see a graphical and table summary of the data as well as a table with the detailed results.