Whole genome alignments, Demo
Let’s look at some of the comparative genomics views in the Location tab. Go to the region 1:8000-18000 in Oryza sativa Japonica.
We can look at individual species comparative genomics tracks in this view by clicking on Configure this page.
Select BLASTz/LASTz alignments from the left-hand menu to choose alignments between closely related species. Turn on the alignments for all Oryza species:

The alignment is greatest between closely related species. We can see that many rice species (such as Oryza barthii) are fully aligned across the region, but other species have a region around 1:10700-12500 where a different chunk is aligned (such as Oryza punctata).
We can also look at the alignment between species or groups of species as text. Click on Alignments (text) in the left hand menu.
Select Select an alignment to open the alignment menu.
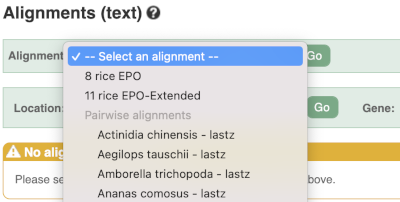
Select Oryza punctata from the alignments list then click Go.
In this case there are eight blocks aligned of different lengths, some of which correspond to the region we saw unaligned in the image. Click on Block 1.
You will see a list of the regions aligned, followed by the sequence alignment. Click on Display full alignment. Exons are shown in red.
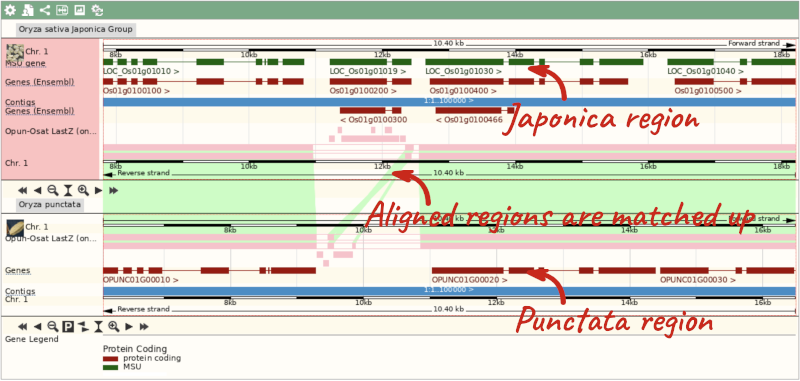
To compare with both contigs visually, go to Region comparison.
To add species to this view, click on the blue Select species or regions button. Choose Oryza punctata again then close the menu.
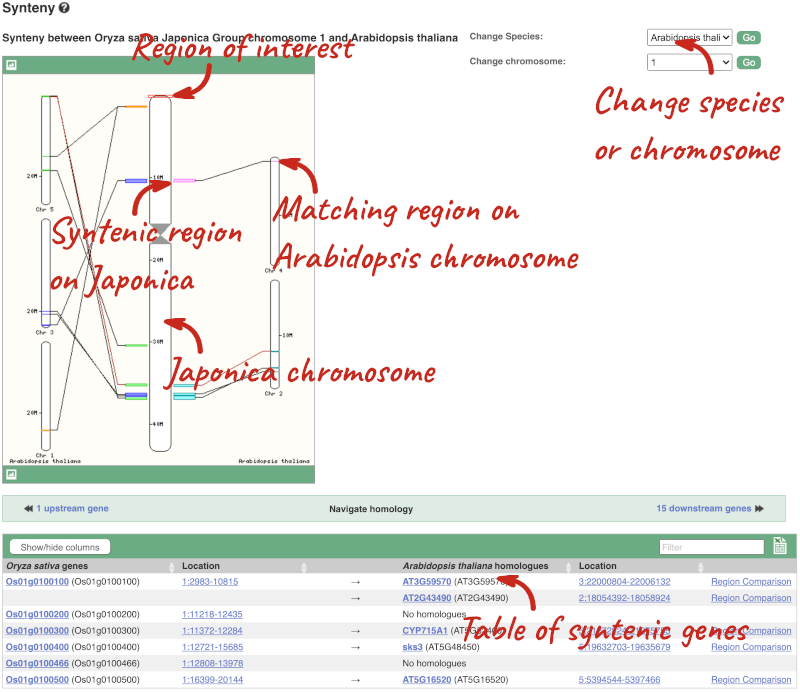
We can view large scale syntenic regions from our chromosome of interest. Click on Synteny in the left hand menu.