Aligning genomic regions, demo
Let’s look at some of the comparative genomics views in the Location tab. Go to the region 15:81873000-82000800 in pig, which contains the HoxD cluster which is involved in limb development and is highly conserved between species.
You can turn on conservation scores and constrained elements. Click on Configure this page, then Comparative genomics and turn on the tracks for Constrained elements for pig breeds and Conservation score for pig breeds. Save and close the menu.
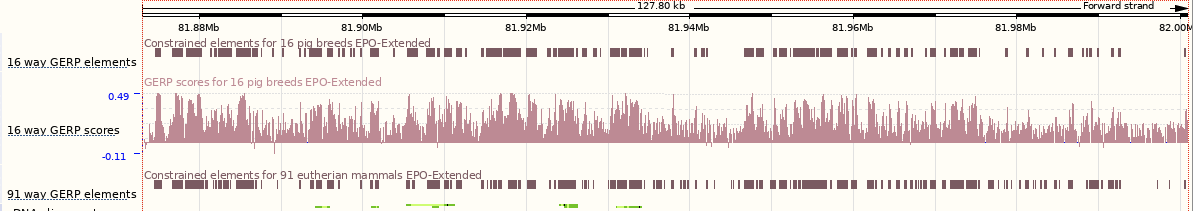
You can now see the conservation scores in pale pink. These were used to determine the peaks indicated in the constrained elements track in dark pink. This track indicates regions of high conservation between species, considered to be “constraine”d by evolution.
We can also look at individual species comparative genomics tracks in this view by clicking on Configure this page.
Select BLASTz/LASTz alignments from the left-hand menu to choose alignments between closely related species. Turn on the alignments for_ Human_ and Cow in Normal. Save and close the menu.

We can also look at the alignment between species or groups of species as text. Click on Alignments (text) in the left hand menu.
Select_ Cow_ from the alignments list then click Go.
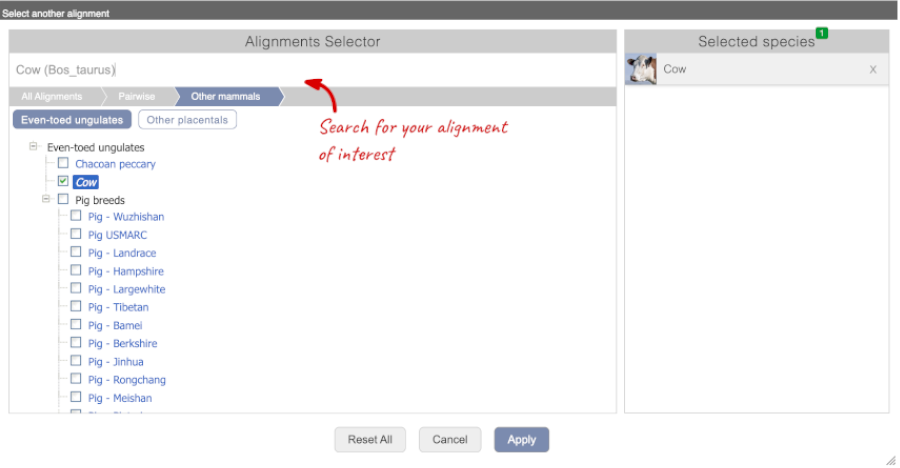
You will see a list of the regions aligned, followed by the sequence alignment. Exons are shown in red.
To compare with both contigs visually, go to Region comparison.
To add species to this view, click on the blue Select species or regions button. Choose Cow from the list then close the menu.
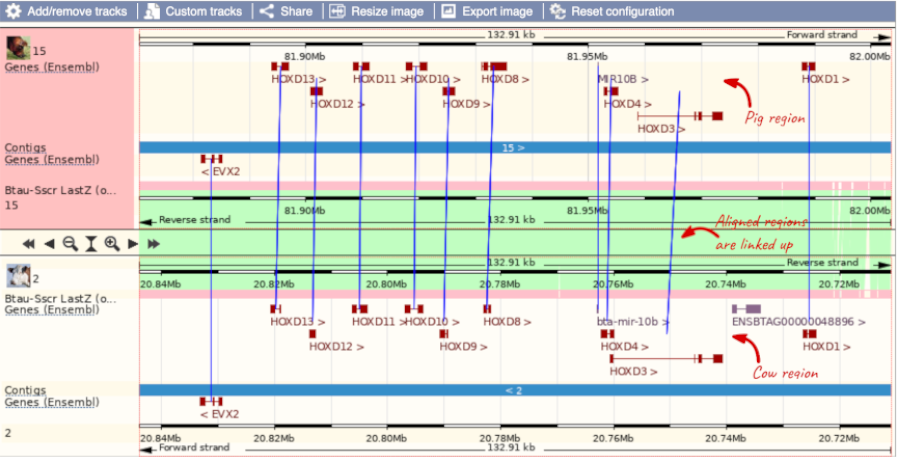
You can configure this view for both species. Click on configure this page and look in the top left of the menu.
The drop down allows you to configure each species separately.
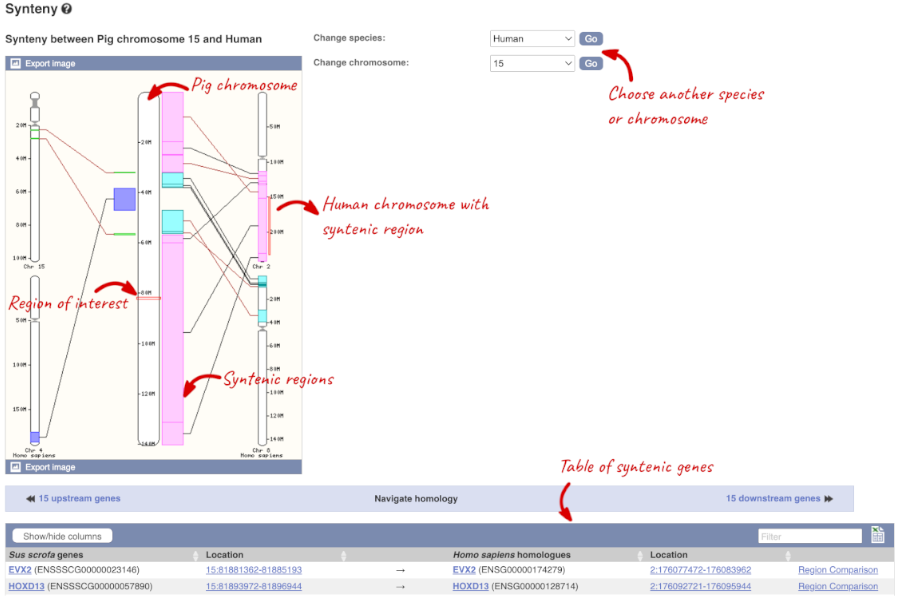
We can view large scale syntenic regions from our chromosome of interest. Click on _Synteny _in the left hand menu.