Gene trees and homologues, Demo
Let’s look at the homologues of human BRCA2. Search for the gene and go to the Gene tab.
Click on Gene tree, which will display the current gene in the context of a phylogenetic tree used to determine orthologues and paralogues.
.png)
Funnels indicate collapsed nodes. We can expand them by clicking on the node and selecting Expand this sub-tree from the pop-up menu.
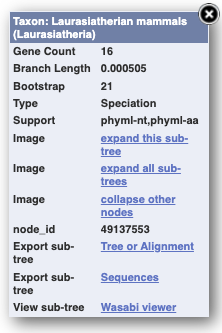
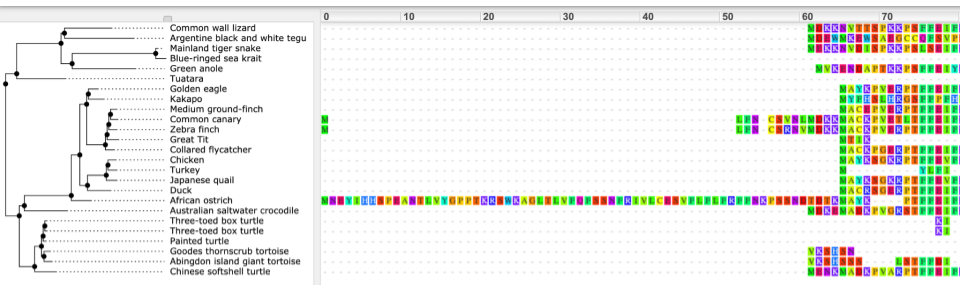
We can also see the alignment of the sub-tree by clicking on Wasabi viewer, which will open a pop-up:

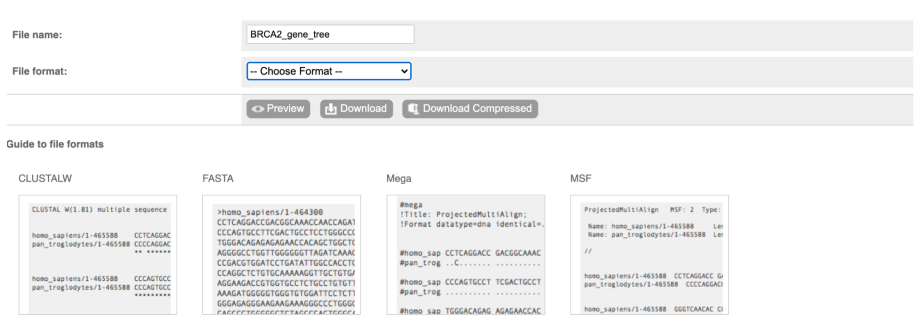
You can download the tree in a variety of formats. Click on the download icon in the bar at the top of the image to get a pop-up where you can choose your format.

We can look at homologues in the Orthologues and Paralogues pages, which can be accessed from the left-hand menu. If there are no orthologues or paralogues, then the name will be greyed out. Paralogues is greyed out for BRCA2 indicating that there are no paralogues available. Click on Orthologues to see the 175 orthologues available.
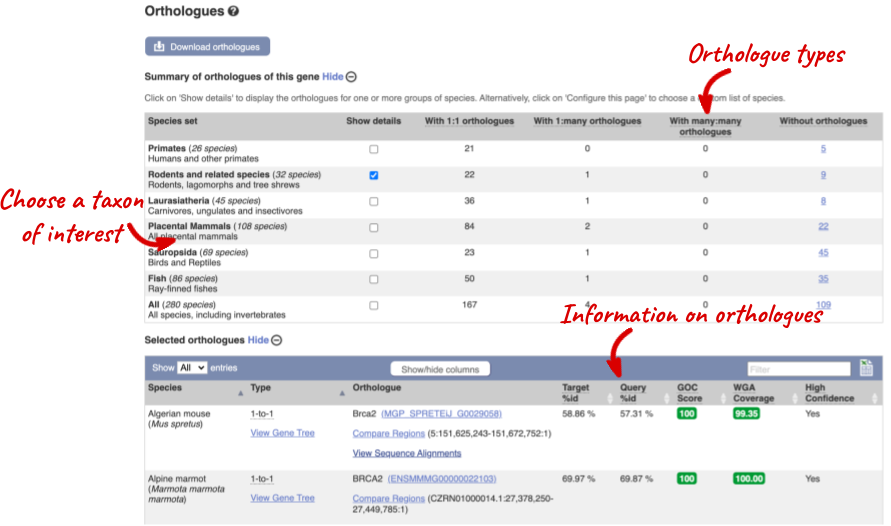
Choose to see only Rodents and related species orthologues by selecting the box. The table below will now only show details of rodent orthologues. Let’s look at mouse.

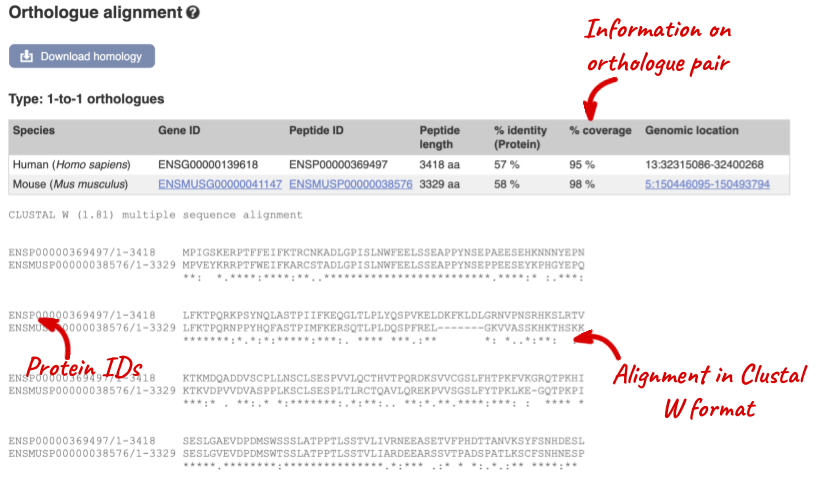
Links from the orthologue allow you to go to alignments of the orthologous proteins and cDNAs. Click on View Sequence Alignments then View Protein Alignment for the mouse orthologue.