Filter Events by Year
Ensembl Variation Workshop - GM7, MSt. in Genomic Medicine, University of Cambridge
Course Details
- Lead Trainer
- Jamie Allen
- Event Date
- 2024-12-04
- Location
- School of Clinical Medicine, University of Cambridge
- Description
- Work with the Ensembl Outreach team to get hands-on experience accessing and analysing variation data with the Ensembl genome browser and the Variant Effect Predictor (VEP).
- Survey
- Ensembl Variation Workshop - GM7, MSt. in Genomic Medicine, University of Cambridge Feedback Survey
Demos and exercises
VEP
We have identified five variants on human chromosome nine, C-> A at 128203516, an A deletion at 128328461, C->A at 128322349, C->G at 128323079 and G->A at 128322917.
We will use the Ensembl VEP to determine:
- Have my variants already been annotated in Ensembl?
- What genes are affected by my variants?
- Do any of my variants affect gene regulation?
Go to the front page of Ensembl and click on the Variant Effect Predictor.

This page contains information about the VEP, including links to download the script version of the tool. Click on Launch VEP to open the input form:
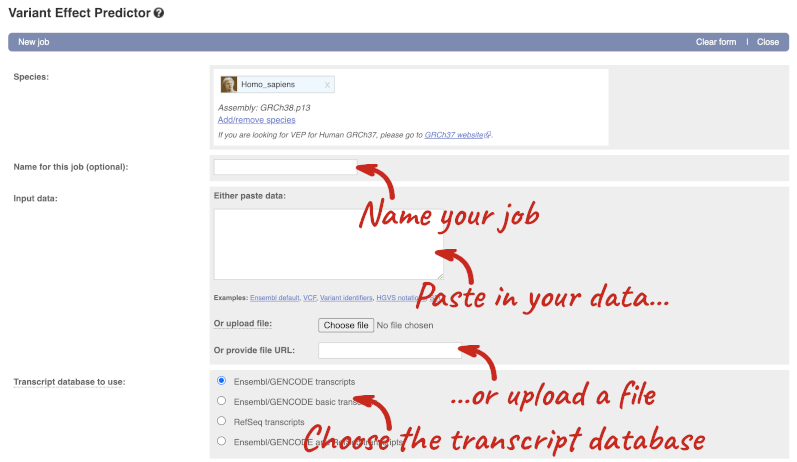
The data is in VCF format:
chromosome coordinate id reference alternative
Put the following into the Paste data box:
9 128328460 var1 TA T
9 128322349 var2 C A
9 128323079 var3 C G
9 128322917 var4 G A
9 128203516 var5 C A
The VEP will automatically detect that the data is in VCF.
There are further options that you can choose for your output. These are categorised as Identifiers, Variants and frequency data, Additional annotations, Predictions, Filtering options and Advanced options. Let’s open all the menus and take a look.
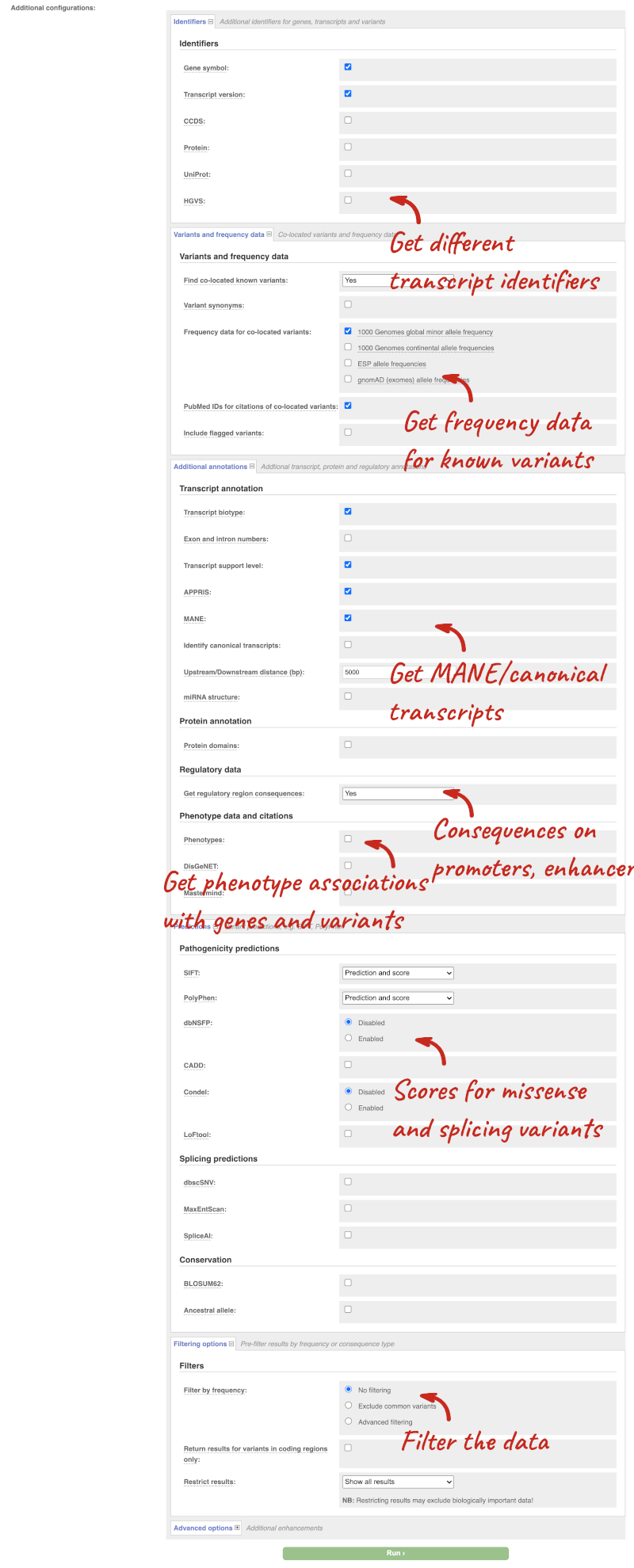
Hover over the options to see definitions.
We’re going to select some options:
- HGVS, annotation of variants in terms of the transcripts and proteins they affect, commonly-used by the clinical community
- Phenotypes
- Protein domains
When you’ve selected everything you need, scroll right to the bottom and click Run.

The display will show you the status of your job. It will say Queued, then automatically switch to Done when the job is done, you do not need to refresh the page. You can edit or discard your job at this time. If you have submitted multiple jobs, they will all appear here.
Click View results once your job is done.
In your results you will see a graphical summary of your data, as well as a table of your results.
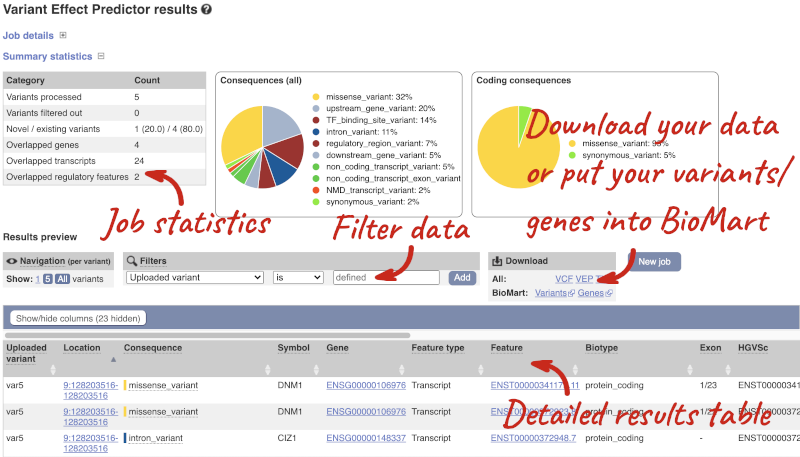
The results table is enormous and detailed, so we’re going to go through the it by section. The first column is Uploaded variant. If your input data contains IDs, like ours does, the ID is listed here. If your input data is only loci, this column will contain the locus and alleles of the variant. You’ll notice that the variants are not neccessarily in the order they were in in your input. You’ll also see that there are multiple lines in the table for each variant, with each line representing one transcript or other feature the variant affects.
You can mouse over any column name to get a definition of what is shown.
The next few columns give the information about the feature the variant affects, including the consequence. Where the feature is a transcript, you will see the gene symbol and stable ID and the transcript stable ID and biotype. Where the feature is a regulatory feature, you will get the stable ID and type. For a transcription factor binding motif (labelled as a MotifFeature) you will see just the ID. Most of the IDs are links to take you to the gene, transcript or regulatory feature homepage.
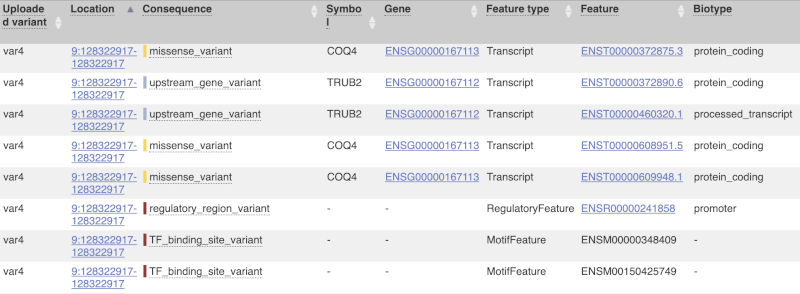
This is followed by details on the effects on transcripts, including the position of the variant in terms of the exon number, cDNA, CDS and protein, the amino acid and codon change, transcript flags, such as MANE, on the transcript, which can be used to choose a single transcript for variant reporting, and predicted pathogenicity scores. The predicted pathogenicity scores are shown as numbers with coloured highlights to indicate the prediction, and you can mouse-over the scores to get the prediction in words. Two options that we selected in the input form are the HGVS notation, which is shown to the left of the image below and can be used for reporting, and the Domains to the right. The Domains list the proteins domains found, and where there is available, provide a link to the 3D protein model which will launch a LiteMol viewer, highlighting the variant position.
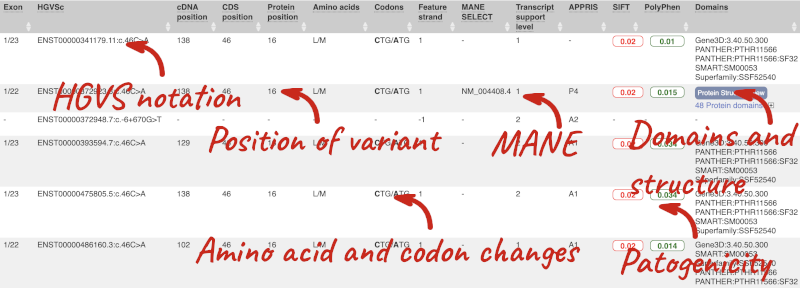
Where the variant is known, the ID of the existing variant is listed, with a link out to the variant homepage. In this example, only rsIDs from dbSNP are shown, but sometimes you will see IDs from other sources such as COSMIC. The VEP also looks up the variant from frequency files, and pulls back the allele frequency (AF in the table), which will give you the 1000 Genomes Global Allele Frequency. In our query, we have not selected allele frequencies from the continental 1000 Genomes populations or from gnomAD, but these could also be shown here. We can also see ClinVar clinical significance and the phenotypes associated with known variants or with the genes affected by the variants, with the variant ID listed for variant associations and the gene ID listed for gene associations, along with the source of the association.
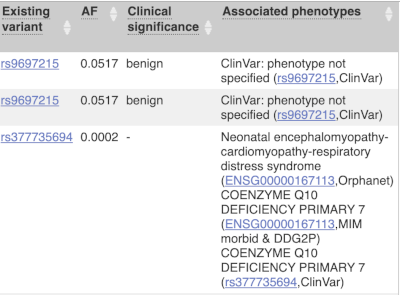
For variants that affect transcription factor binding motifs, there are columns that show the effect on motifs (you may need to click on Show/hide columns at the top left of the table to display these). Here you can see the position of the variant in the motif, if the change increases or decreases the binding affinity of the motif and the transcription factors that bind the motif.
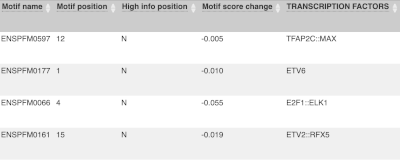
Above the table is the Filter option, which allows you to filter by any column in the table. You can select a column from the drop-down, then a logic option from the next drop-down, then type in your filter to the following box. We’ll try a filter of Consequence, followed by is then missense_variant, which will give us only variants that change the amino acid sequence of the protein. You’ll notice that as you type missense_variant, the VEP will make suggestions for an autocomplete.
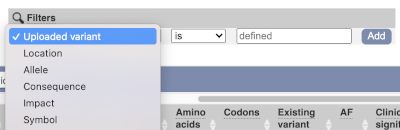
You can export your VEP results in various formats, including VCF. When you export as VCF, you’ll get all the VEP annotation listed under CSQ in the INFO column. After filtering your data, you’ll see that you have the option to export only the filtered data. You can also drop all the genes you’ve found into the Gene BioMart, or all the known variants into the Variation BioMart to export further information about them.
Running CFTR variants through VEP
Resequencing of the genomic region of the human CFTR (cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) gene (ENSG00000001626) has revealed the following variants. The alleles defined in the forward strand:
- G/A at 7: 117,530,985
- T/C at 7: 117,531,038
- T/C at 7: 117,531,068
Use the VEP tool in Ensembl and choose the options to see SIFT and PolyPhen predictions. Do these variants result in a change in the proteins encoded by any of the Ensembl genes? Which gene? Have the variants already been found?
Go to the Ensembl homepage and click on the link Tools at the top of the page. Currently there are nine tools listed in that page. Click on Variant Effect Predictor and enter the three variants as below:
7 117530985 117530985 G/A
7 117531038 117531038 T/C
7 117531068 117531068 T/C
Note: Variation data input can be done in a variety of formats. See more details about the different data formats and their structure in this VEP documentation page. Click Run. When your job is listed as Done, click View Results.
You will get a table with the consequence terms from the Sequence Ontology project (http://www.sequenceontology.org/) (i.e. synonymous, missense, downstream, intronic, 5’ UTR, 3’ UTR, etc) provided by VEP for the listed SNPs. You can also upload the VEP results as a track and view them on Location pages in Ensembl. SIFT and PolyPhen are available for missense SNPs only. For two of the entered positions, the variations have been predicted to have missense consequences of various pathogenicity (coordinate 117531038 and 117531068), both affecting CFTR. All the three variants have been already annotated and are known as rs1800077, rs1800078 and rs35516286 in dbSNP (databases, literature, etc).
VEP analysis of structural variants in human
We have details of a genomic deletion in a breast cancer sample in VCF format:
13 32307062 sv1 . <DEL> . . SVTYPE=DEL;END=32908738
Use VEP in Ensembl to find out the following information:
1. How many genes have been affected?
2. Does the structural variant (SV) cause deletion of any complete transcripts?
3. Map your variant in the Ensembl browser on the Region in detail view.
- Click on VEP at the top of any Ensembl page and open the web interface. Make sure your species is Human. It is good practise to name your VEP jobs something descriptive, such as Patient deletion exercise. Paste the variant in VCF format into the Paste data field and hit Run.
12 different genes are affected by the SV.
- Filter your table by select Consequence is
transcript_ablationat the top of the table and click Add.Yes, there is deletion of complete transcripts of PDS5B, N4BP2L1, BRCA2, RNY1P4, IFIT1P1, ATP8A2P2, N4BP2L2, N4BP2L2-IT2 and one gene without official symbols: ENSG00000212293.
- To view your variant in the browser click on the location link in the results table 13: 32307062-32908738. The link will open the Region in detail view in a new tab. If you have given your data a name it will appear automatically in red. If not, you may need to Configure this page and add it under the Personal data tab in the pop-up menu.
VEP CMD Human
Command-line VEP analysis of variants from a 1000 Genomes Project dataset
Ensembl’s Variant Effect Predictor (VEP) is a powerful tool for annotating genomic variants. Ensembl VEP is accessible via web, REST API and command line options.
In this practical session, we will practice using VEP via the command line to annotate a variant call file (VCF).
The VCF you will use contains variant calls for Homo Sapiens chromosome 13 from the IGSR: The International Genome Sample Resource. This file was extracted using Ensembl’s Data Slicer for human variation data and is aligned to GRCh38, with the IGSR British in England and Scotland GBR population samples subsetted. The file is available here
Directories and starting tutorials:
A General Tutorial for command line VEP is available to try out or compare to if you need some guidance.
You can use the options guide at Ensembl VEP Options to help find the commands you need. TIP: this is very useful for finding the commands you will need!
Launch a command prompt. VEP is installed already and should work directly from the terminal command line. You can either run in the location you are when you open the terminal, or create a directory for this work (something like mkdir vep_practical).
The subsetted IGSR VCF file name is: 13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf and you can download it by running:
wget https://ftp.ebi.ac.uk/pub/databases/ensembl/training/output_files/13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf
Exercises:
Ex 1. Explore the VCF file with any text viewer to explore its contents. What do lines denoted by “##” represent? What are the key headers in the file?
You can use VCF file specifications as a reminder.
Ex 2. Use the command-line VEP tool to annotate the variants in the IGSR VCF file and output to VCF format. Here is an example command:
vep -i [input_file] -o [output_file] --cache --offline
We recommend using the default VEP output format, as it’s easier to read on screen, but you can preserve the output as VCF by using --vcf.
Ex 3. Explore the output of VEP in any text viewer to explore the contents. What genes are affected by the variants in the file? Why are there multiple results for each variant?
The VEP cache contains Allele Frequency (AF) data available for known variants from multiple sources. As this data is from the 1000 genomes it already contains allele frequency data from that resource.
Ex 4. On the original input file (13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf), rerun Ensembl VEP, this time use the appropriate options to add AF data from gnomAD genomes, gene symbols, and to identify MANE Select transcripts. Write the output to a new file (something like -o chr13_with_AF_symbols_mane.txt).
Ex 5. Use the filter_vep tool on the output of the previous step to find variants that are:
-
in
BRCA2and -
are a
MANE_SELECTtranscript and -
which have an AF of 0.01 or less in the overall gnomAD genomes population
gnomADg_AF. Careful with this one - if a variant doesn’t have any AF data it will be filtered out as not matching the gnomAD annotation - HINT: consider using not.
Why do you think we have used an AF filter here?
Ex 6. From the original output of exercise 2, what has been the overall effect of adding annotations and then filtering using that information? Are there any missense variants left? How might you filter for just missense variants?
Bonus!
If you finish all that, try pasting the variant below into Web VEP and select the alphamissense, revel, CADD, mavedb and mutfunc options.
What do you see in the results?
13 48040982 . A T
Exercise 1
You can open the VCF file with any text editor, or by using a command such as: less -S 13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf. Then use the arrow keys to navigate, and press Q when ready to quit. Lines starting with ## indicate information headers, which contain information about the file content and how it was processed.
Exercise 2
vep -i 13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf -o VEP_annot_chr13_GBR.vcf --cache --offline
Your own script may not look exactly like this and you may employ different flags, here are some common examples:
--input_fileor-iAllows you to specify the location of the input file.--output_fileor-oAllows you to specify the name of the output file.--force_overwriteAllows VEP to overwrite a pre-existing output file with the same name.--genomesPoints VEP to the Ensembl Genomes (non-vertebrates) server. (not used for this homo sapiens example)--cacheEnables the use of the cache (this can speed up VEP significantly).--cache_versonAllows you to specify the cache version. This should be used with Ensembl Genomes caches, since their version numbers do not match Ensembl versions. For example, the VEP/Ensembl version may be 110 and the Ensembl Genomes version 57.--check_existingChecks for the existence of known variants that are co-located with your input variants.--offlineEnables offline mode (no database connections are made).
Exercise 3
You may use a similar less command or method to open the annotated VCF file:
less -S VEP_annot_chr13_GBR.vcf
There are two genes listed in the annotations, ZAR1L ENSG00000189167 and BRCA2 ENSG00000139618. There are multiple rows for each variant representing the variant consequence in different transcripts.
Exercise 4
Rerun VEP with the following command:
vep -i 13.32315086-32400268.ALL.chr13_GRCh38.genotypes.20170504.GBR.vcf -o VEP_annot_chr13_GBR_MANE_SYMB_gnomadAF.txt --mane --cache --symbol --af_gnomadg --offline
Exercise 5
Filter for BRCA2 and MANE with the following command:
filter_vep -i VEP_annot_chr13_GBR_MANE_SYMB_gnomadAF.txt -o VEP_annot_FILT.txt --filter "((SYMBOL is BRCA2) and (MANE_SELECT exists) and (gnomADg_AF <= 0.01 or not gnomADg_AF))"
Exercise 6
By adding MANE transcript annotation, gene symbol and allele frequency information, then using filters based on these, we can reduce the annotated variant set down from 1770 to 81.
To filter to include only the missense results, for example, you could add and (Consequence is missense_variant) to the command above.
There is a missense variant - rs80358762 - you can explore phenotype information related to that variant here. Note that although it’s a rare missense variant in BRCA2, the evidence from multiple ClinVar submitters indicate that it is benign.
Bonus!
This variant is novel and does not have an accessioned ID and is not present in the allele frequency resources. Consider just the missense variant in NUDT15, you can see that prediction tools we enabled (alphamissense, revel and CADD) are all relatively low scores, below those that would normally be thought of as pathogenic. However, some of the mutfunc and MaveDB scores - which are based on experimental data - indicate that the protein structure is significantly impacted, suggesting this variant may merit further investigation.







