Filter Events by Year
ESHG: Using the Ensembl Variant Effect Predictor (VEP)
Course Details
- Lead Trainer
- Aleena Mushtaq
- Event Dates
- 2024-06-01 until 2024-06-04
- Location
- Berlin
- Description
- This workshop will introduce participants to how the Ensembl Variant Effect Predictor can be used to interpret genetic variation in clinical research in the context of Ensembl/GENCODE annotation and MANE transcripts
Demos and exercises
Genes and variants
Ensembl homepage
The front page of Ensembl is found at ensembl.org. It contains lots of information and links to help you navigate Ensembl:
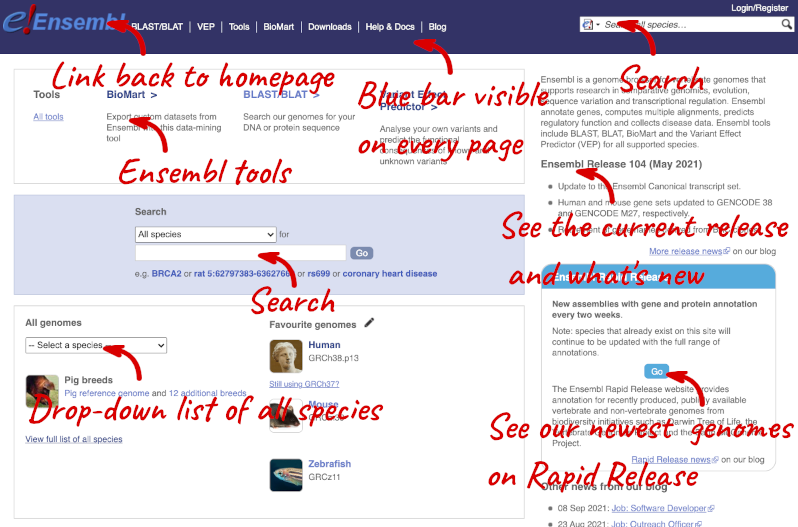
At the top left you can see the current release number and what has come out in this release. To access old releases, scroll to the bottom of the page and click on View in archive site.
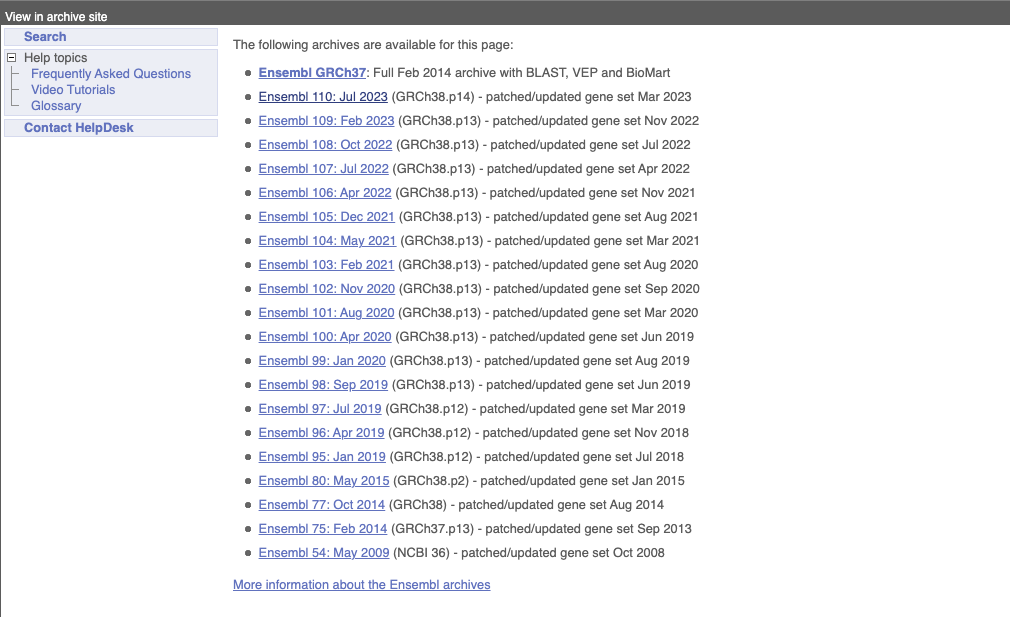
Click on the links to go to the archives. Alternatively, you can jump quickly to the correct release by putting it into the URL, for example e98.ensembl.org jumps to release 98.
Click on View full list of all species.
Click on the common name of your species of interest to go to the species homepage. We’ll click on Human.
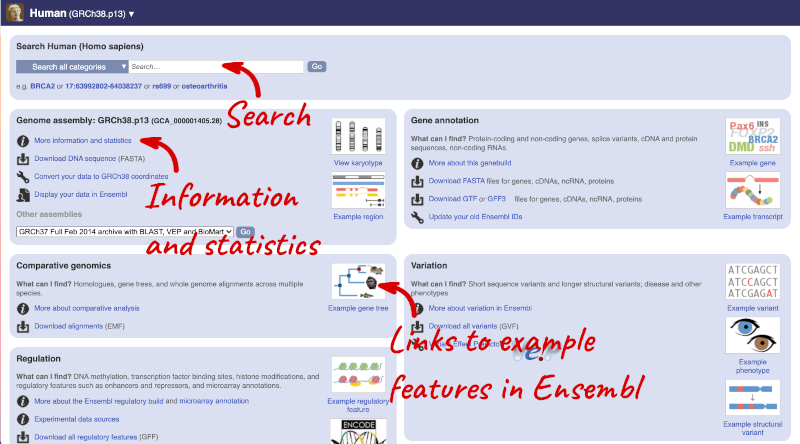
Here you can see links to example pages and to download flatfiles. To find out more about the genome assembly and genebuild, click on More information and statistics.
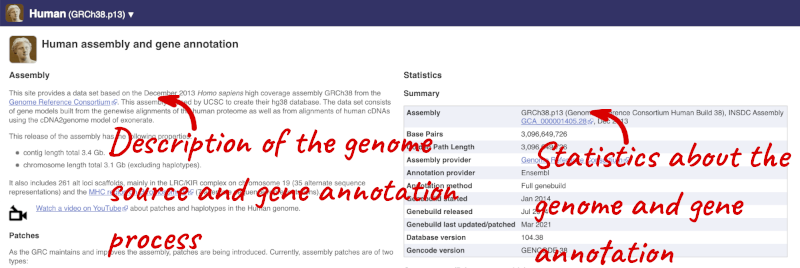
Here you’ll find a detailed description of how to the genome was produced and links to the original source. You will also see details of how the genes were annotated.
The current genome assembly for human is GRCh38. If you want to see the previous assembly, GRCh37, visit our dedicated site grch37.ensembl.org.
The gene tab
You can find out lots of information about Ensembl genes and transcripts using the browser. You can search for gene names or identifiers, and also phenotypes or functions that might be associated with the genes.
We’re going to look at the human JAK2 gene. From ensembl.org, type JAK2 into the search bar and click the Go button. You will get a list of hits with the human gene at the top.
Where you search for something without specifying the species, or where the ID is not restricted to a single species, the most popular species will appear first, in this case, human, mouse and zebrafish appear first. You can restrict your query to species or features of interest using the options on the left.
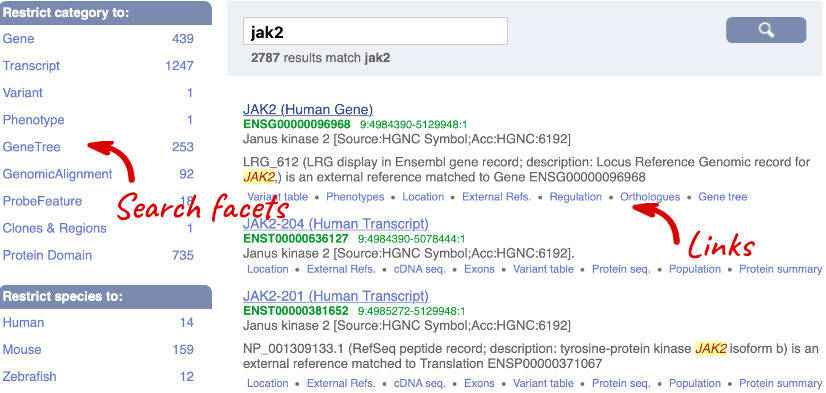
Click on the gene name or Ensembl ID. The Gene tab should open:
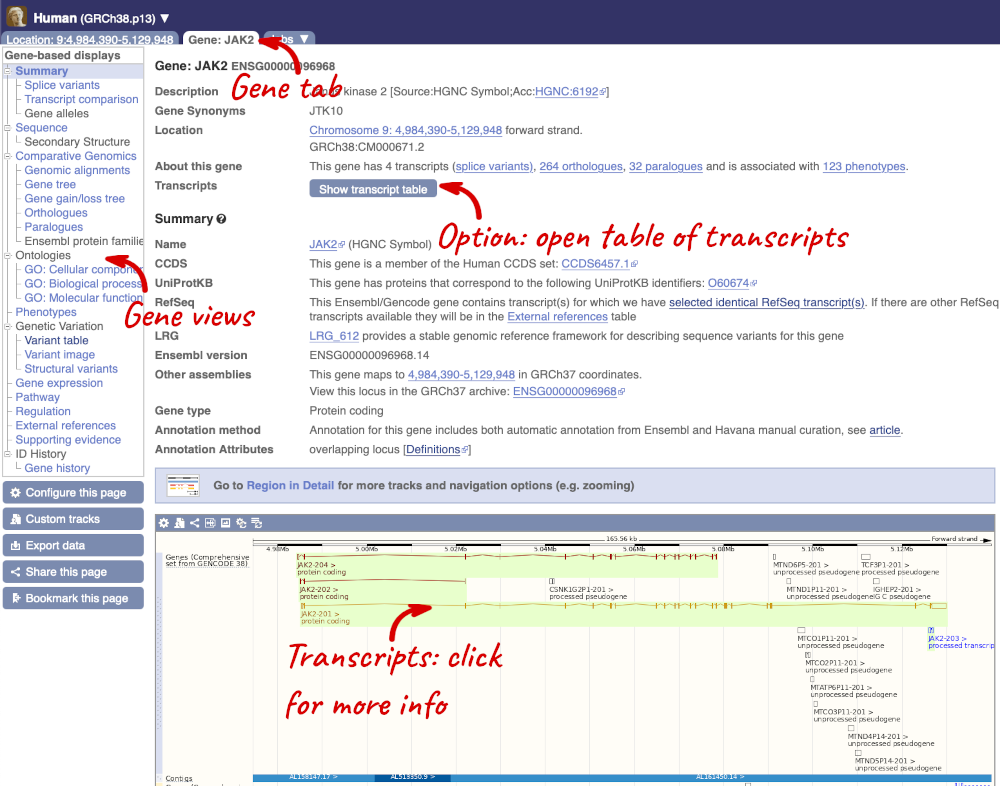
This page summarises the gene, including its location, name and equivalents in other databases. At the bottom of the page, a graphic shows a region view with the transcripts. We can see exons shown as blocks with introns as lines linking them together. Coding exons are filled, whereas non-coding exons are empty. We can also see the overlapping and neighbouring genes and other genomic features.
There are different tabs for different types of features, such as genes, transcripts or variants. These appear side-by-side across the blue bar, allowing you to jump back and forth between features of interest. Each tab has its own navigation column down the left hand side of the page, listing all the things you can see for this feature.
Let’s walk through this menu for the gene tab. How can we view the genomic sequence? Click Sequence at the left of the page.
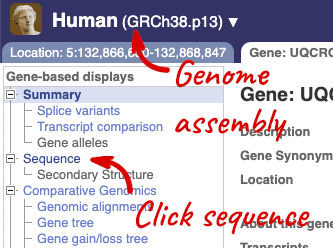
The sequence is shown in FASTA format. The FASTA header contains the genome assembly, chromosome, coordinates and strand (1 or -1) – this gene is on the positive strand.

Exons are highlighted within the genomic sequence, both exons of our gene of interest and any neighbouring or overlapping gene. By default, 600 bases are shown up and downstream of the gene. We can make changes to how this sequence appears with the blue Configure this page button found at the left. This allows us to change the flanking regions, add variants, add line numbering and more. Click on it now.
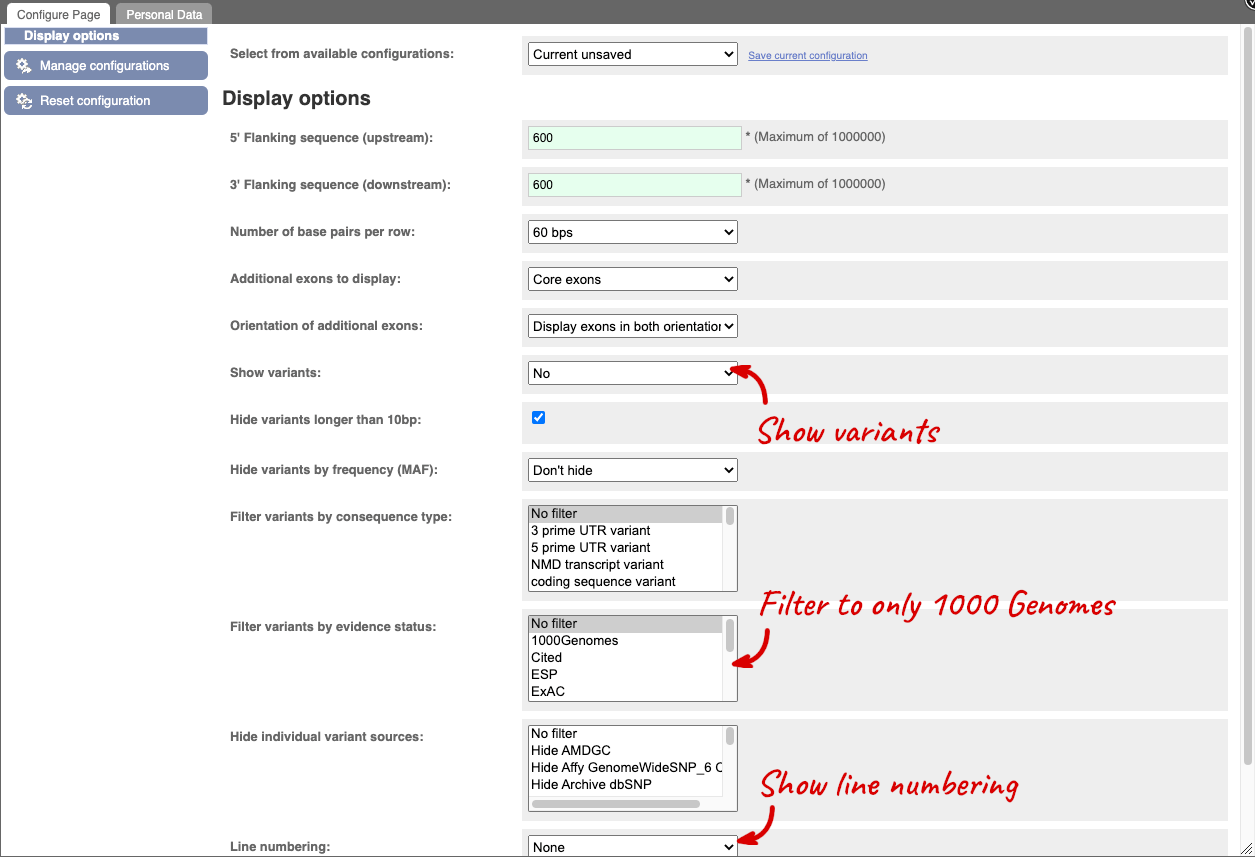
You can view variants on the sequence in any of the sequence views. Once you have selected changes (in this example, Show variants: Yes and show links, Filter variants by evidence status: 1000Genomes and Line numbering: Relative to coordinate system) click at the top right.
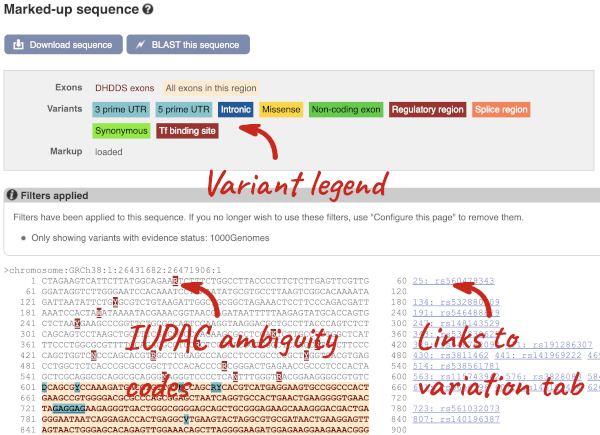
Find out more about a variant by clicking on it.
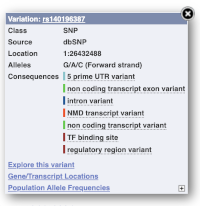
You can go to the Variation tab by clicking on the variant ID. For now, we’ll explore more ways of finding variants.
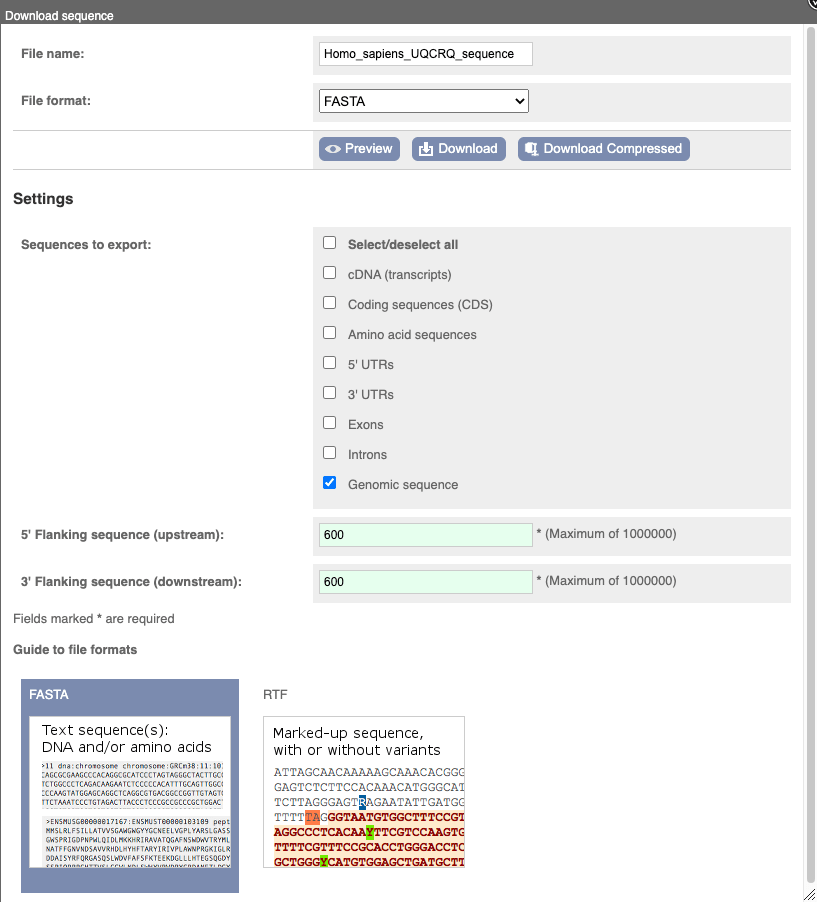
You can download this sequence by clicking in the Download sequence button above the sequence:

This will open a dialogue box that allows you to pick between plain FASTA sequence, or sequence in RTF, which includes all the coloured annotations and can be opened in a word processor. If you want run a sequence analysis tool, download as FASTA sequence, whereas if you want to analyse the sequence visually, RTF is best for this. This button is available for all sequence views.
To view all the sequence variants in table form, click the Variant table link at the left of the gene tab.
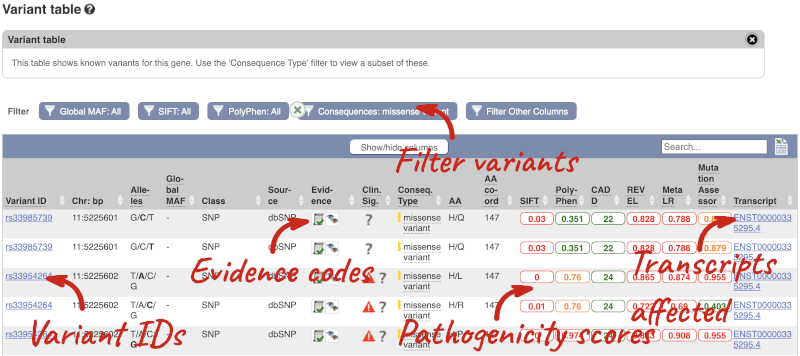
You can filter the table to only show the variants you’re interested in. For example, click on Consequences: All, then select the variant consequences you’re interested in. For display purposes, the table above has already been filtered to only show missense variants.
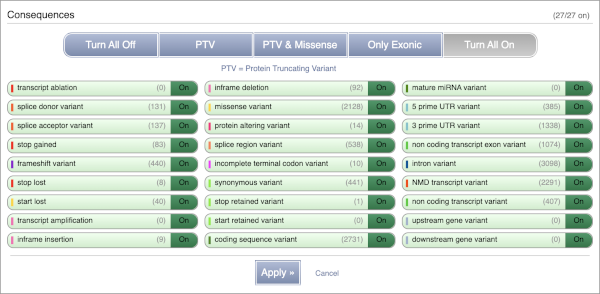
You can also filter by the different pathogenicity scores and MAF, or click on Filter other columns for filtering by other columns such as Evidence or Class.
The table contains lots of information about the variants. You can click on the IDs here to go to the Variation tab too.
You can also see the phenotypes associated with a gene. Click on Phenotype in the left hand menu.
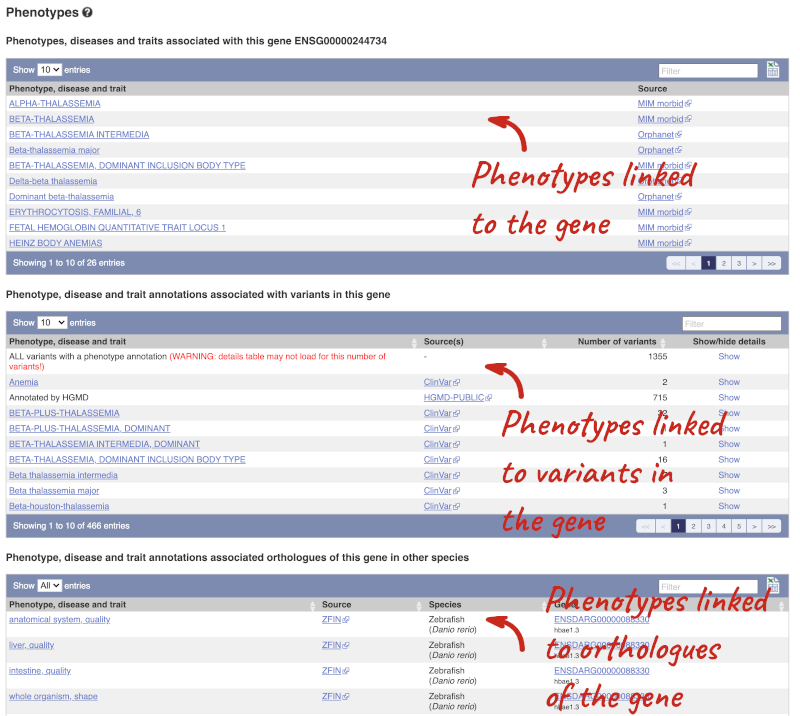
The transcript tab
We’re now going to explore the different transcripts of JAK2. Click on Show transcript table at the top.

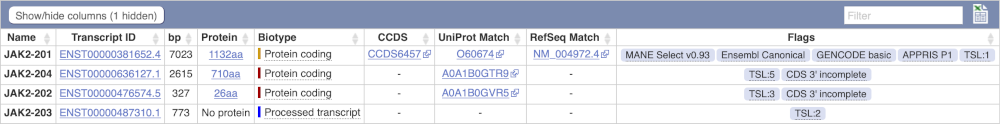
Here we can see a list of all the transcripts of JAK2 with their identifiers, lengths, biotypes and flags to help you decide which ones to look at.
If we were to only choose one transcript to analyse, we would choose JAK2-201 because it is the MANE Select and Ensembl Canonical. This means it is both 100% identical to the RefSeq transcript NM_004972.4 and both Ensembl and NCBI agree that it is the most biologically important transcript.
Click on the ID, ENST00000381652.4.
You are now in the Transcript tab for JAK2-201. We can still see the gene tab so we can easily jump back. The left hand navigation column provides several options for the transcript JAK2-201 - many of these are similar to the options you see in the gene tab, but not all of them. If you can’t find the thing you’re looking for, often the solution is to switch tabs.
Click on the Exons link. This page is useful for designing RT-PCR primers because you can see the sequences of the different exons and their lengths.
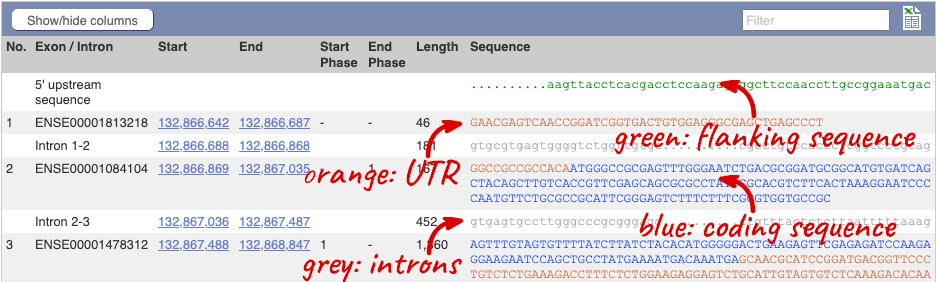
You may want to change the display (for example, to show more flanking sequence, or to show full introns). In order to do so click on Configure this page and change the display options accordingly.
Now click on the cDNA link to see the spliced transcript sequence with the amino acid sequence. This page is useful for mapping between the RNA and protein sequences, particularly genetic variants.
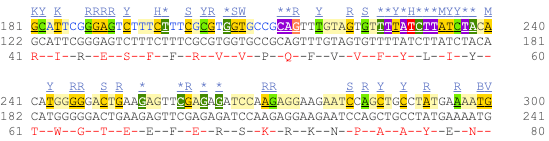
UnTranslated Regions (UTRs) are highlighted in dark yellow, codons are highlighted in light yellow, and exon sequence is shown in black or blue letters to show exon divides. Sequence variants are represented by highlighted nucleotides and clickable IUPAC codes are above the sequence.
Click on Haplotypes in the left hand menu.

The Haplotypes view in the transcript tab shows you the actual protein and CDS sequences in 1000 Genomes individuals. This is possible because the 1000 Genomes study has phased genotypes, so we know which alleles occur on which of the chromosome pairs. The table lists all the versions of the protein that occur along with their frequencies, including the reference sequence and sequences with one or more alternative alleles.
Click on one of the haplotypes, we’ll go for 393L>V, to find out more about it. Here you will see the frequency in the 1000 Genomes subpopulations, the sequence and the 1000 Genomes individuals where this protein is found.
The variant tab
Blood cancers known as MPNs commonly occur due to an activating somatic mutation (JAK2-V617F) in the JAK2 gene decades before diagnosis. Let’s have a look at a specific variant. We could find the variant COSV67569051 in the Variant table, however it’s easier to find if we put COSV67569051 into the search box. Click through to open the Variation tab.
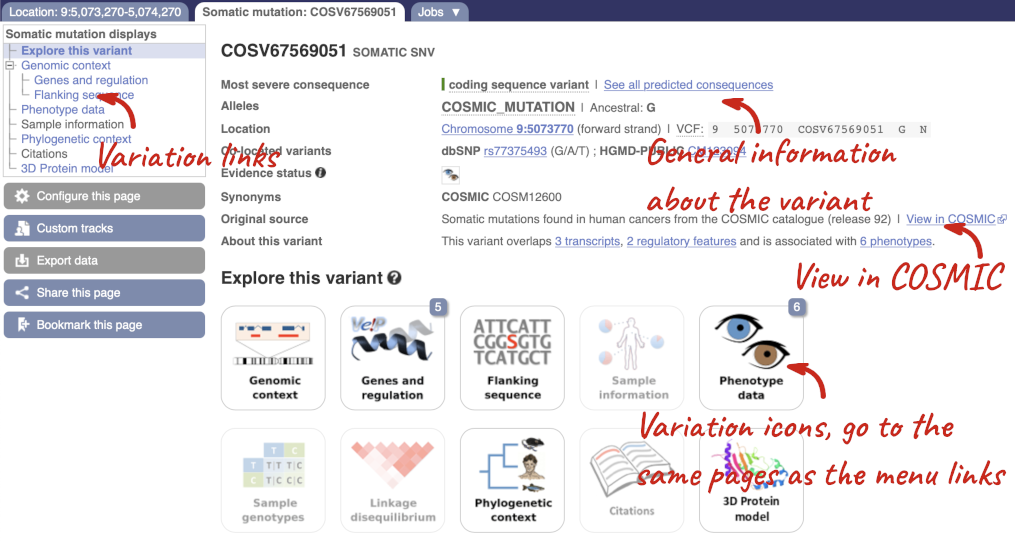
The icons show you what information is available for this variant. Click on Genes and regulation, or follow the link on the left.
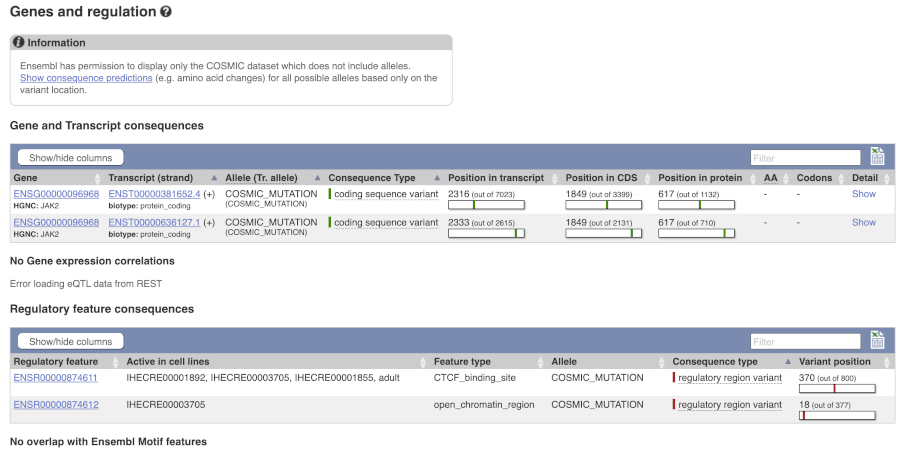
We can see the variant falls in a coding region, as the consequence is coding sequence variant. Ensembl cannot display the alleles for COSMIC variants, so you will need go to the COSMIC database to find out if it is a missense or synonymous mutation. Click on View in COSMIC in the panel above, and then c.1849G>T to go to the variant page.
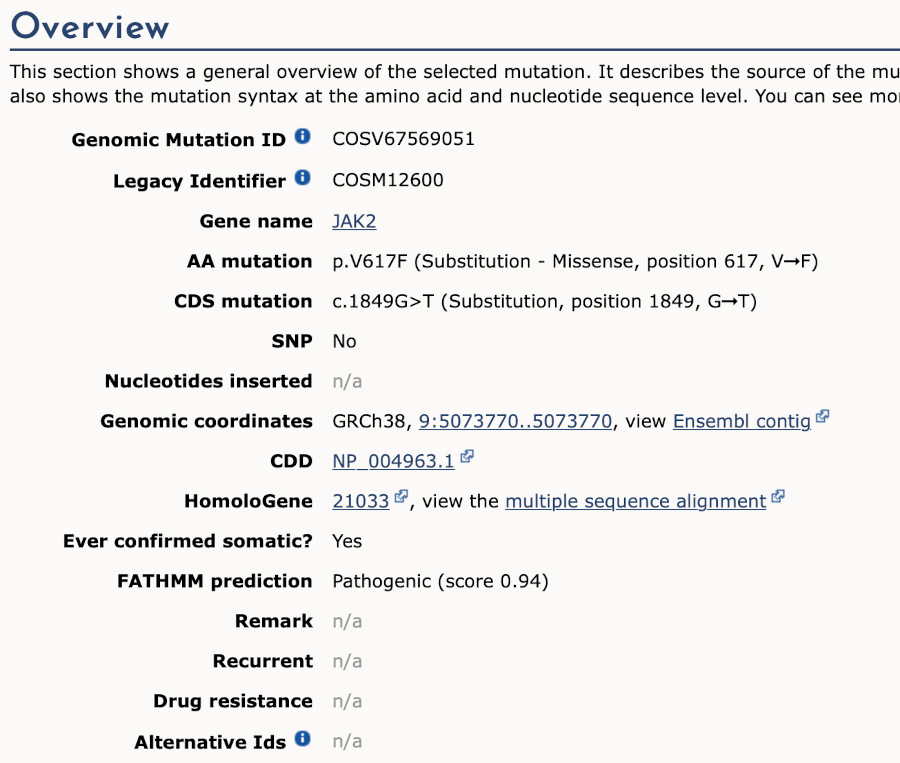
We now know that this variant alleles are G>T, causing V617F amino acid substitution. Go back to the variant tab in Ensembl and click on dbSNP variant rs77375493 under Co-located variants in the overview panel.

It is the same variant, but imported from dbSNP. We can see that there is more information for this dbSNP variant, compared to the COSMIC variant. We can see the consequence, alternate alleles and allele frequency data. Let’s click on Genes and regulation now.
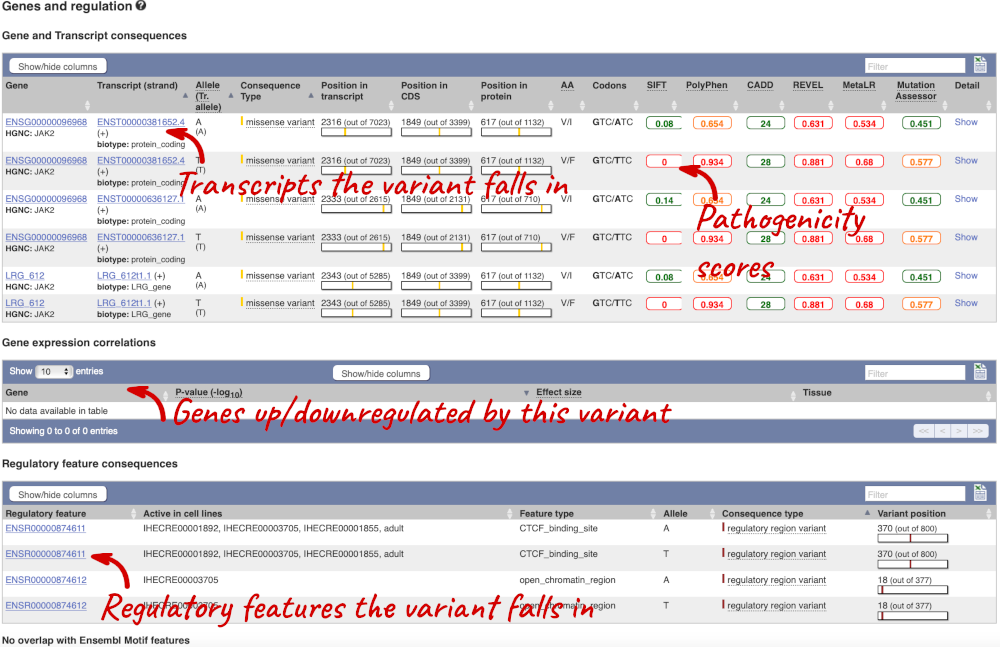
This page illustrates the genes the variant falls within and the consequences on those genes, including pathogenicity predictors, but also regulatory features that the variant falls within.
Let’s look at population genetics. Click on Population genetics in the left-hand menu.
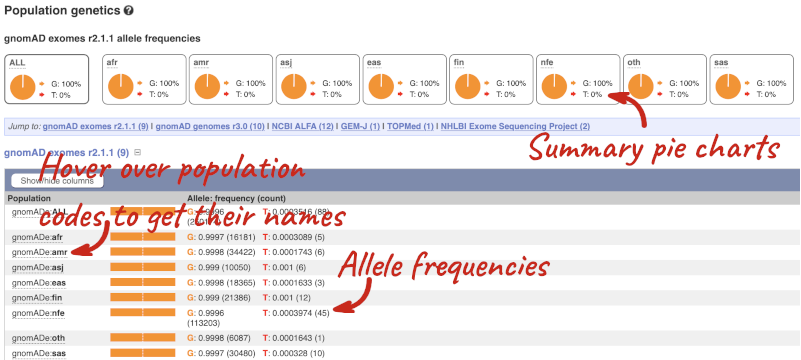
This variant is very rare and has not been found in 1000 Genomes populations, but has been reported in some diseased cohorts. The population allele frequencies are shown by study, including gnomAD, NCBI ALPHA, GEM-J, TOPMed and NHLBI Exome Sequencing Project. Where genotype frequencies are available, these are shown in the tables.
Let’s have a look at the phenotypes associated with this variant. Click on Phenotype Data in the left-hand menu.
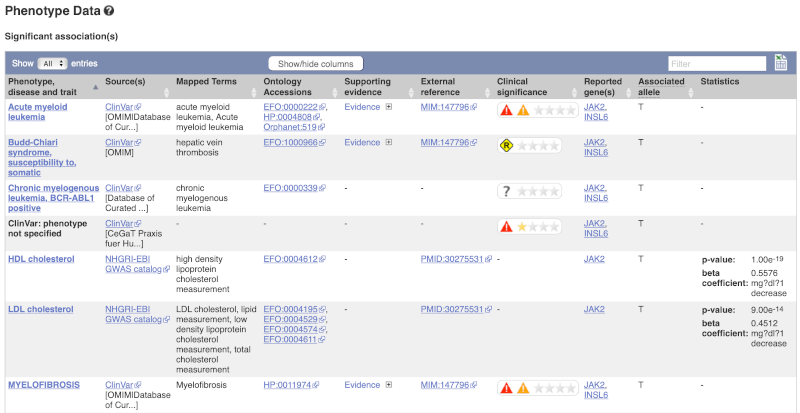
This variant is associated with various phenotypes, including blood cancers such as acute myeloid leukemia, myelofibrosis and increased numbers of mature blood cells. These phenotype associations come from sources including the GWAS catalog, ClinVar, Orphanet and OMIM. Where available, the table provides links to the original paper that made the association, the allele that is associated with the phenotype and p-values and other statistics.
Are there other variants in the genome that also cause acute myeloid leukemia? Click on the phenotype Acute myeloid leukemia to find out.
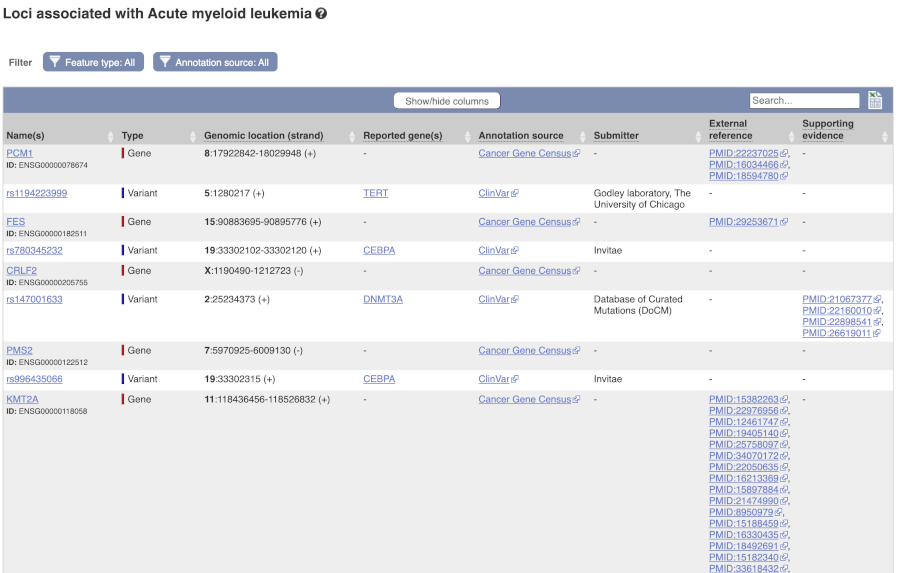
Human population genetics and phenotype data
The SNP rs1738074 in the 5’ UTR of the human TAGAP gene has been identified as a genetic risk factor for a few diseases. Use Ensembl to answer the following questions:
-
In which transcripts is this SNP found?
-
What is the least frequent genotype for this SNP in the Yoruba (YRI) population from the 1000 Genomes phase 3?
-
What is the ancestral allele? Is it conserved in the 91 eutherian mammals EPO-Extended?
-
With which diseases is this SNP associated? Are there any known risk (or associated) alleles?
- Please note there is more than one way to get this answer. Either go to the Variation table of the human TAGAP gene, and use the Consequence filter to only include 5’UTR variants, or search Ensembl for
rs1738074directly. Once you’re in the Variant tab, click on Genes and regulation in the menu.This SNP is found in four transcripts of TAGAP. It is also an intron_variant to one lncRNA transcript of TAGAP-AS1.
- Click on Population genetics in the left-hand panel, or click on Explore this variant in the left-hand panel and click the Population genetics icon.
In Yoruba (YRI), the least frequent genotype is CC at the frequency of 5.6%.
- Click on Phylogenetic context in the left-hand panel.
The ancestral allele is T and it’s inferred from the alignment in primates.
Click on Select an alignment which will open a pop-up menu. Open Multiple alignments and select 91 eutherian mammals EPO-Extended. Click on Apply at the bottom of the menu to save your settings.
A region containing the SNP (highlighted in red and placed in the centre) and its flanking sequence are displayed. The T allele is conserved in all but two of the eutherian mammals displayed.
- Click Phenotype data in the left-hand panel.
This variation is associated with multiple sclerosis, celiac disease and white blood cell count. There are known risk alleles for all three diseases and the corresponding P values are provided. The allele A is associated with celiac disease. Note that the alleles reported by Ensembl are T/C. Ensembl reports alleles on the forward strand. This suggests that A was reported on the reverse strand in the original paper. Similarly, one of the alleles reported for Multiple sclerosis is G.
Exploring a SNP in the human genome
The missense variation rs1801133 in the human MTHFR gene has been linked to elevated levels of homocysteine, an amino acid whose plasma concentration seems to be associated with the risk of cardiovascular diseases, neural tube defects, and loss of cognitive function.
-
Find the page with information for rs1801133.
-
Is rs1801133 a missense variant in all transcripts of the MTHFR gene? What is the amino acid change?
-
Why are the alleles for this variation in Ensembl given as G/A and not as C/T, as in the literature?
-
What is the major allele of rs1801133 in different populations?
-
In which paper(s) is the association between rs1801133 and homocysteine levels described?
-
According to the data imported from dbSNP, the ancestral allele for rs1801133 is G. Ancestral alleles in dbSNP are based on a comparison between human and chimp. Does the sequence at this same position in other primates confirm that the ancestral allele is G?
-
Go to the Ensembl homepage. Type
rs1801133in the search box, then click Go. Click on rs1801133. -
Click on Genes and Regulation in the left-hand panel, or click on the Genes and Regulation icon at the top of the page.
No, rs1801133 is missense variant in eight MTHFR transcripts. Please note that this variant is multialleleic with two alternative alleles - as this table displays one consequence per row, each transcript is listed twice.
The amino acid change is A/V for allele A, and A/G for allele C.
-
In Ensembl, the alleles of rs1801133 are given as G/A/C because these are the alleles in the forward strand of the genome. In the literature, the alleles are given as C/T/G because the MTHFR gene is located on the reverse strand. The alleles in the actual gene and transcript sequences are C/T/G. In Ensembl, the allele that is present in the reference genome assembly is always put first.
- Click on Population genetics in the side menu.
In all populations but one, the allele G is the major one. The exception is CLM (Colombian in Medellin; 1000 Genomes).
- Click on Phenotype Data in the left hand side menu.
The specific studies where the association was originally described is given in the Phenotype Data table. Links between rs1801133 and homocysteine levels were described in four papers. Click on the pubmed IDs PMID:34707639, PMID:23696881, PMID:20031578 and PMID:23824729 for more details.
- Click on Phylogenetic Context in the side menu. Select Alignment: 10 primates EPO and click Apply.
Gorilla, bonobo, Sumatran orangutan, chimp, macaque, gibbon, vervet, crab-eating macaque and mouse lemur all have a G in this position.
Exploring the MYH9 gene in human
- In Ensembl, find the human MYH9 (myosin, heavy chain 9, non-muscle) gene and open the Gene tab.
- On which chromosome and which strand of the genome is this gene located?
- How many transcripts (splice variants) are there and how many are protein coding?
- What is the longest protein-coding transcript, and how long is the protein it encodes?
- Which transcript would you take forward for further study?
-
Click on Phenotypes at the left side of the page. Are there any diseases associated with this gene, according to Mendelian Inheritance in Man (MIM)?
-
What are some functions of MYH9 according to the Gene Ontology (GO) consortium? Have a look at the GO: Biological process pages for this gene.
- In the transcript table, click on the transcript ID for MYH9-201, and go to the Transcript tab.
- How many exons does it have?
- Are any of the exons completely or partially untranslated?
- Is there an associated sequence in UniProtKB/Swiss-Prot? Have a look at the General identifiers for this transcript.
- Are there microarray (oligo) probes that can be used to monitor ENST00000216181 expression?
- Select Human from the Species drop-down list and type
MYH9. Click Go. Click on MYH9 (Human Gene) in the search results which will send you to the Gene tab.- The gene is located on chromosome 22 on the reverse strand.
- Ensembl has 23 transcripts annotated for this gene, of which 6 are protein-coding.
- The longest protein-coding transcript is MYH9-215 and it codes for a protein that is 1,981 amino acids long.
- MYH9-201 is the transcript I would take forward for further study, as it is the MANE Select transcript (for a description, mouse-over the MANE Select flag in the transcript table).
- Click on Phenotypes in the left-hand panel to see the associated phenotypes. There is a large table of phenotypes. To see only the ones from MIM, type
MIMinto the filter box at the top right-hand corner of the table.These are some of the phenotypes associated with MYH9 according to MIM: Deafness, Autosomal dominant 17 and Macrothrombocytopenia and granulocyte inclusions with or without nephritis or sensorineural hearing loss. You can click on the records for more information.
-
The Gene Ontology project maps terms to a protein in three classes: biological process, cellular component, and molecular function. Click on GO: Biological process on the left-hand panel. Angiogenesis, cell adhesion, and protein transport are some of the roles associated with MYH9. All GO terms are associated with a single transcript: ENST00000216181.
- Click on ENST00000216181.11 in the transcript table. You should now be on the Transcript tab.
- It has 41 exons, shown in the Transcript summary.
Click on the Exons link in the left-hand panel.
- Exon 1 is completely untranslated, and exons 2 and 41 are partially untranslated (UTR sequence is shown in orange). You can also see this in the cDNA view if you click on the cDNA link in the left side menu.
Click on General identifiers in the left-hand panel.
- P35579.254 from UniProt/Swiss-Prot matches the translation of the Ensembl transcript. Click on P35579.254 to go to UniProtKB, or click align for the alignment.
- Click on Oligo probes in the left-hand panel.
Probesets from Affymetrix, Agilent, Codelink, Illumina, and Phalanx OneArray match to this transcript sequence. Expression analysis with any of these probesets would reveal information about the transcript. Hint: this information can sometimes be found in the [ArrayExpress Atlas] (https://www.ebi.ac.uk/biostudies/arrayexpress).
Finding a gene associated with a phenotype
Phenylketonuria is a genetic disorder caused by an inability to metabolise phenylalanine in any body tissue. This results in an accumulation of phenylalanine causing seizures and intellectual disability.
(a) Search for phenylketonuria from the Ensembl homepage and narrow down your search to only genes. What gene is associated with this disorder?
(b) How many protein coding transcripts does this gene have? View all of these in the transcript comparison view.
(c) What is the MIM gene identifier for this gene?
(d) Go to the MANE Select transcript and look at its 3D structure. In the model 2pah, how many protein molecules can you see?
(a) Start at the Ensembl homepage (http://www.ensembl.org).
Type phenylketonuria into the search box then click Go. Choose Gene from the left hand menu.
The gene associated with this disorder is PAH, phenylalanine hydroxylase, ENSG00000171759.
(b) If the transcript table is hidden, click on Show transcript table to see it.
There are six protein coding transcripts.
Click on Transcript comparison in the left hand menu. Click on Select transcripts. Either select all the transcripts labelled protein coding one-by-one, or click on the drop down and select Protein coding. Close the menu.
(c) Click on External references.
The MIM gene ID is 612349.
(d) Open the transcript table and click on the ID for the MANE Select: ENST00000553106.6. Go to PDB 3D protein model in the left-hand menu.
The model 2pah is shown by default. It has two protein molecules in it. You may need to rotate the model to see this clearly.
Exploring VNTR in human
Variable number tandem repeats (VNTRs) show high variation in the number of repeats in the population and are commonly used in forensics (DNA fingerprinting) and to study genetic diversity. (a) Go to the region from 3074666 to 3075100 bp on human chromosome 4. Which gene does it overlap? Which exon of this gene falls in this region?
(b) Configure this page to turn on Repeats (low), Simple repeats (Repeats (low)) and Tandem repeats (TRF) tracks in this view. Can you see any repeats in this exon? What tools were used to annotate the repeats according to the track information?
(c) Zoom in on the (CAG)n to see its sequence. How many CAG repeats can you see in the human reference assembly? Does this track overlap any phenotype-associated variants? What is the identifier of this variant?
(d) Go to the variant tab of the phenotype-associated variant. What is the consequence ontology of this variant? Does the reference allele match the number of repeats you have just counted? What is the shortest and longest allele?
(a) Select Search: Human and type 4:3074666-3075100 in the text box (or alternatively type human 4:3074666-3075100 in the text box). Click Go.
Click on the golden transcript falling in this region. You can see it’s exon 1 of 67 of the huntingtin gene (HTT).
(b) Click Configure this page in the side menu then select: Repeats (low), Simple repeats (Repeats (low)) and Tandem repeats (TRF).
There are three tandem repeats in this exon, and two simple repeats (low); (CAG)n and (CCG)n. Click on the track names to find more about the tools used for annotation: RepeatMasker and Tandem Repeats Finder.
(c) Draw with your mouse a box around the (CAG)n repeat. Click on Jump to region in the pop-up menu.
There are 19 CAG repeats in the human reference sequence overlapping rs71180116 indicated by a pink bar in the All phenotype-associated - short variants (SNPs and indels) track.
(d) Click on the rs71180116 ID to go to the variant tab. You can see in the summary page that this variant is classified as an inframe insertion. Either click + to show all of the alleles in the summary page or go to the Genes and regulation table. This variant has many alternative alleles which differ in the number of repeats. The first allele in the expanded Alleles section of the summary page or the first allele in the Codons column in the Genes and regulation table is the reference allele. It is composed of 19 CAG repeats just as in the Region in detail view. The shortest allele has 7 repeats, the longest has 55 repeats.
VEP
We have identified 10 variants on human chromosome eight in a patient with neurodegenerative symptoms including dystonia.
We will use the Ensembl VEP to determine:
- Have my variants already been annotated in Ensembl?
- What genes are affected by my variants?
- Do any of my variants affect gene regulation?
Go to the front page of Ensembl and click on the Variant Effect Predictor.

This page contains information about the VEP, including links to download the script version of the tool. Click on Launch VEP to open the input form:
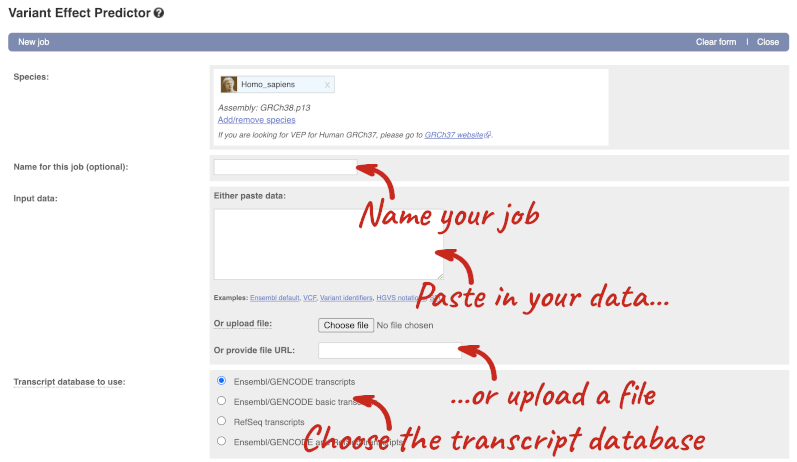
The data is in VCF format:
chromosome coordinate id reference alternative
Put the following into the Paste data box:
8 22410000 var1 G A
8 22412087 var2 C G
8 22408305 var3 T C
8 22409953 var4 C T
8 22414115 var5 C T
8 22406904 var6 GA C
8 22366942 var7 G A
8 22408348 var8 C T
8 22404716 var9 GCTGCTGCTGCTGC GCTGCTGC
8 22404808 var10 T C
The VEP will automatically detect that the data is in VCF.
There are further options that you can choose for your output. These are categorised as Identifiers, Variants and frequency data, Additional annotations, Predictions, Filtering options and Advanced options. Let’s open all the menus and take a look.
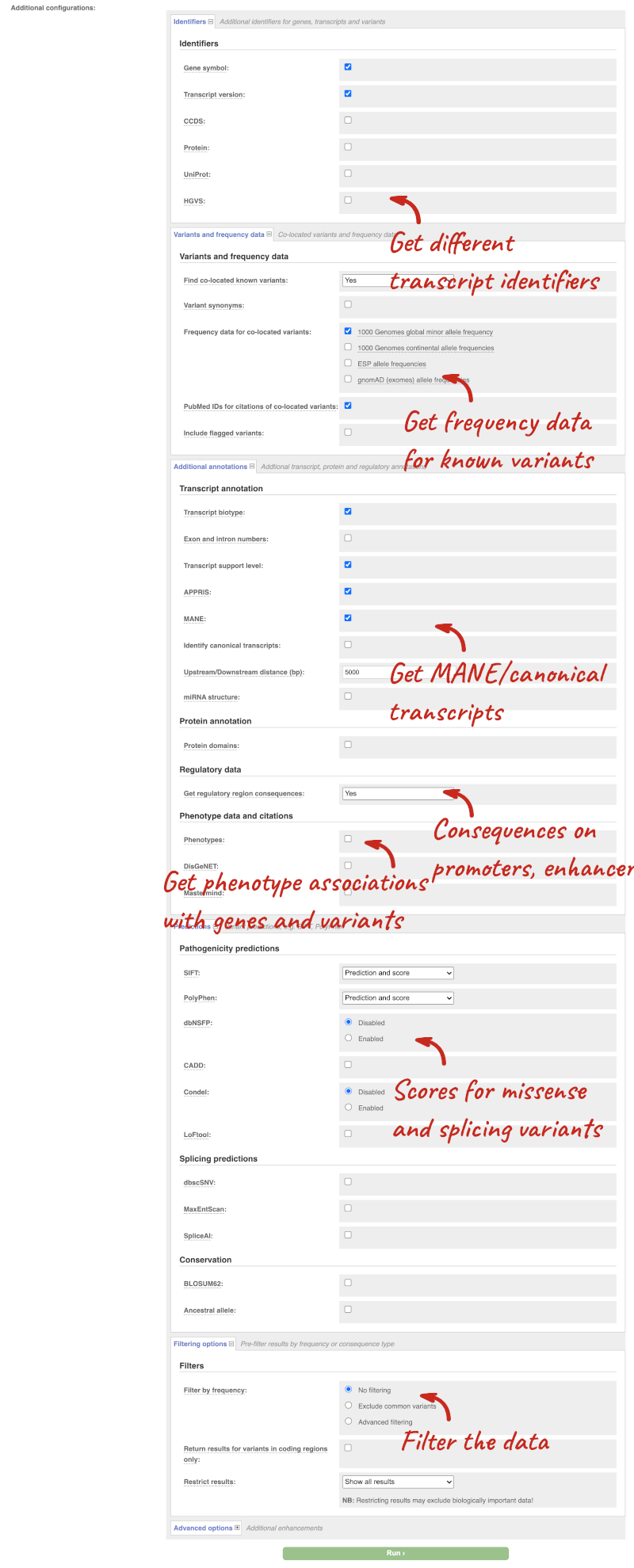
Hover over the options to see definitions.
We’re going to select some options:
- HGVS, annotation of variants in terms of the transcripts and proteins they affect, commonly-used by the clinical community
- gnomAD (exomes) allele frequencies
- Protein matches
- Phenotypes
When you have selected everything you need, scroll right to the bottom and click Run.

The display will show you the status of your job. It will say Queued, then automatically switch to Done when the job is done, you do not need to refresh the page. You can edit or discard your job at this time. If you have submitted multiple jobs, they will all appear here.
Click View results once your job is done.
In your results you will see a graphical summary of your data, as well as a table of your results.
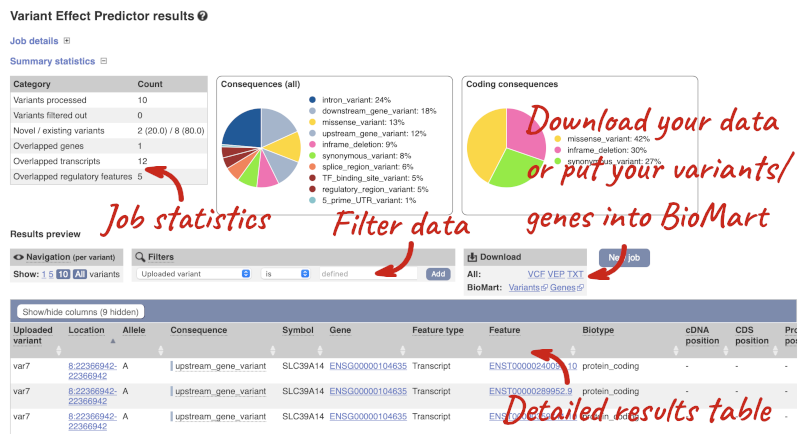
The results table is enormous and detailed, so we’re going to go through it by section. The first column is Uploaded variant. If your input data contains IDs, like ours does, the ID is listed here. If your input data is only loci, this column will contain the locus and alleles of the variant. You will notice that the variants are not neccessarily in the order they were in your input. You will also see that there are multiple lines in the table for each variant, with each line representing one transcript or other feature the variant affects.
You can mouse-over any column name to get a definition of what is shown.
The next few columns give the information about the feature the variant affects, including the consequence. Where the feature is a transcript, you will see the gene symbol and stable ID and the transcript stable ID and biotype. Where the feature is a regulatory feature, you will get the stable ID and type. For a transcription factor binding motif (labelled as a Motif name) you will see just the ID. Most of the IDs are links to take you to the gene, transcript or regulatory feature homepage.
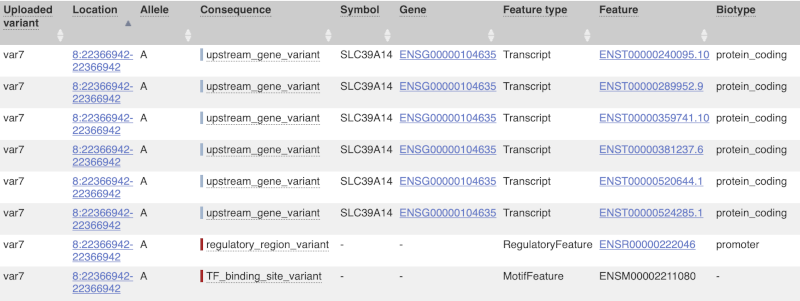

This is followed by details on the effects on transcripts, including the position of the variant in terms of the exon number, cDNA, CDS and protein, the amino acid and codon change, transcript flags, such as MANE, on the transcript, which can be used to choose a single transcript for variant reporting, and pathogenicity scores. The pathogenicity scores are shown as numbers with coloured highlights to indicate the prediction, and you can mouse-over the scores to get the prediction in words. Two options that we selected in the input form are the HGVS notation, which is shown to the left of the image below and can be used for reporting, and the Protein matches to the right. The Protein matches list the proteins domains found, and where available, provide a link to the 3D protein model which will launch a LiteMol viewer, highlighting the variant position.
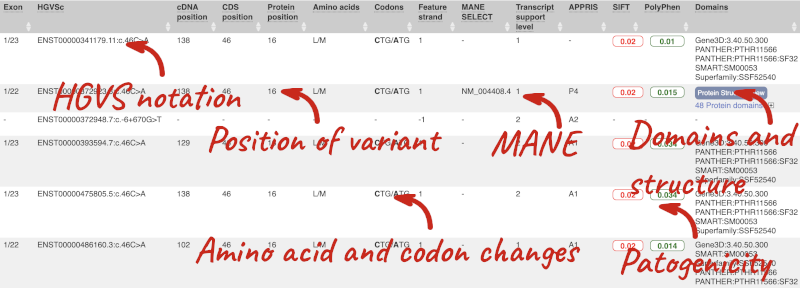
Where the variant is known, the ID of the existing variant is listed, with a link out to the variant homepage. In this example, IDs from dbSNP and COSMIC are shown, but sometimes you will see IDs from other sources as well. The VEP also looks up the variant in the Ensembl database and pulls back the allele frequency (AF in the table), which will give you the 1000 Genomes Global Allele Frequency and allele frequencies from gnomAD. We can also see ClinVar clinical significance and the phenotypes associated with known variants or with the genes affected by the variants, with the variant ID listed for variant associations and the gene ID listed for gene associations, along with the source of the association.
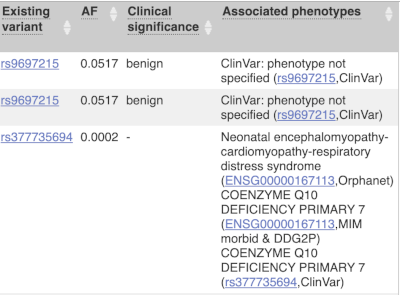
For variants that affect transcription factor binding motifs, there are columns that show the effect on motifs (you may need to click on Show/hide columns at the top left of the table to display these). Here you can see the position of the variant in the motif, if the change increases or decreases the binding affinity of the motif and the transcription factors that bind the motif.
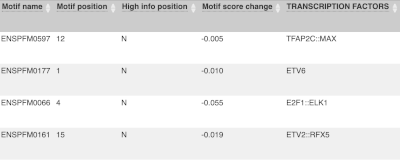
Let’s sort the table by Impact using the little upward triangle next to the column name (you may need to click on Show/hide columns to find it).
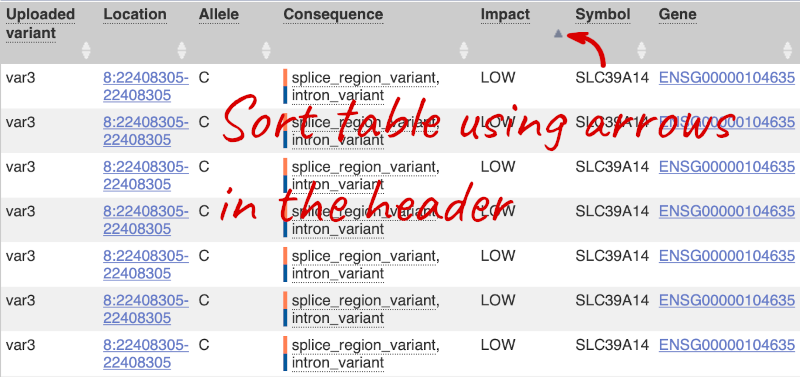
Impact is a subjective classification of the severity of the variant consequence, based on agreement with other variant annotators. It groups variant consequences into four categories: high, moderate, low and modifier. There is no high impact variant on our list, but we have some moderate variant classes such as missense or inframe deletion. Let’s filter the table to get rid of low impact variants now.
Above the table is the Filter option, which allows you to filter by any column in the table. You can select a column from the drop-down, then a logic option from the next drop-down, then type in your filter in the following box. We will try a filter of Impact, followed by is then moderate, which will give us only variant consequences, which might change protein effectiveness. You’ll notice that as you type moderate, the VEP will make suggestions for an autocomplete.
We are still left with quite a few variants. As we are dealing with a rare disorder, we can filter them by the allele frequency assuming that varaints causal of rare diseases should be very rare in the population. Choose gnomADe AF as the filter now, followed by < then 0.005, which will leave us with rare variants found below 0.5% frequency in the general population.
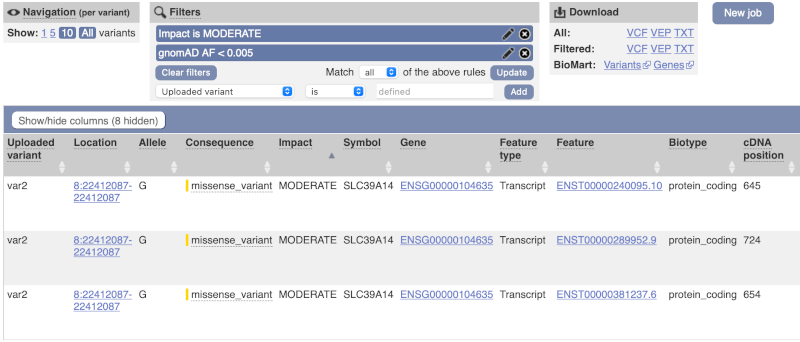
Only one variant meets our criteria. It is a benign missense variant reported on three different transcripts of the SLC39A14 gene, including the MANE Select transcript (NM_001128431.4). Note that the allele frequency for some rare or novel variants might be unknown. Let’s take that into account by editing our allele frequency filter. Change the gnomADe AF filter to is not then defined (leave the text box empty to set it to default defined).
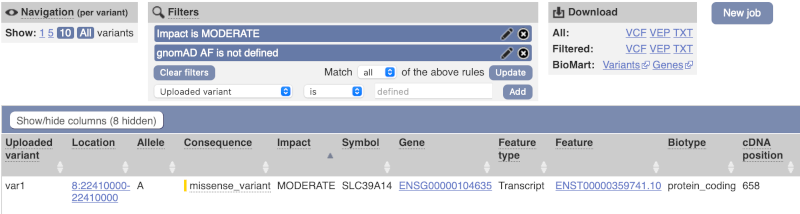
We are left with one pathogenic missense variant of unknown frequency reported on the MANE Plus clinical transcript (NM_015359.6) and associated with hypermagnesemia with dystonia, which can potentially explain our clinical case.
You can export your VEP results in various formats, including VCF. When you export as VCF, you will get all the VEP annotation listed under CSQ in the INFO column. After filtering your data, you will see that you have the option to export only the filtered data. You can also drop all the genes you have found into the Gene BioMart, or all the known variants into the Variation BioMart to export further information about them.
Running CFTR variants through VEP
Resequencing of the genomic region of the human CFTR (cystic fibrosis transmembrane conductance regulator (ATP-binding cassette sub-family C, member 7) gene (ENSG00000001626) has revealed the following variants. The alleles defined in the forward strand:
- G/A at 7: 117,530,985
- T/C at 7: 117,531,038
- T/C at 7: 117,531,068
Use the VEP tool in Ensembl and choose the options to see SIFT and PolyPhen predictions. Do these variants result in a change in the proteins encoded by any of the Ensembl genes? Which gene? Have the variants already been found?
Go to the Ensembl homepage and click on the link Tools at the top of the page. Currently there are nine tools listed in that page. Click on Variant Effect Predictor and enter the three variants as below:
7 117530985 117530985 G/A
7 117531038 117531038 T/C
7 117531068 117531068 T/C
Note: Variation data input can be done in a variety of formats. See more details about the different data formats and their structure in this VEP documentation page. Click Run. When your job is listed as Done, click View Results.
You will get a table with the consequence terms from the Sequence Ontology project (http://www.sequenceontology.org/) (i.e. synonymous, missense, downstream, intronic, 5’ UTR, 3’ UTR, etc) provided by VEP for the listed SNPs. You can also upload the VEP results as a track and view them on Location pages in Ensembl. SIFT and PolyPhen are available for missense SNPs only. For two of the entered positions, the variations have been predicted to have missense consequences of various pathogenicity (coordinate 117531038 and 117531068), both affecting CFTR. All the three variants have been already annotated and are known as rs1800077, rs1800078 and rs35516286 in dbSNP (databases, literature, etc).
VEP analysis of structural variants in human
We have details of a genomic deletion in a breast cancer sample in VCF format:
13 32307062 sv1 . <DEL> . . SVTYPE=DEL;END=32908738
Use VEP in Ensembl to find out the following information:
1. How many genes have been affected?
2. Does the structural variant (SV) cause deletion of any complete transcripts?
3. Map your variant in the Ensembl browser on the Region in detail view.
- Click on VEP at the top of any Ensembl page and open the web interface. Make sure your species is Human. It is good practise to name your VEP jobs something descriptive, such as Patient deletion exercise. Paste the variant in VCF format into the Paste data field and hit Run.
12 different genes are affected by the SV.
- Filter your table by select Consequence is
transcript_ablationat the top of the table and click Add.Yes, there is deletion of complete transcripts of PDS5B, N4BP2L1, BRCA2, RNY1P4, IFIT1P1, ATP8A2P2, N4BP2L2, N4BP2L2-IT2 and one gene without official symbols: ENSG00000212293.
- To view your variant in the browser click on the location link in the results table 13: 32307062-32908738. The link will open the Region in detail view in a new tab. If you have given your data a name it will appear automatically in red. If not, you may need to Configure this page and add it under the Personal data tab in the pop-up menu.







