Filter Events by Year
Ensembl Browser Workshop - Livestock genomics
Course Details
- Lead Trainer
- Aleena Mushtaq
- Event Dates
- 2024-03-18 until 2024-03-21
- Location
- Virtual
- Description
- Work with the Ensembl Outreach team to get to grips with the Ensembl browser, accessing gene, variation, comparative genomics data in common livestock species, and mine these data with BioMart.
Demos and exercises
Ensembl species
The front page of Ensembl is found at ensembl.org. It contains lots of information and links to help you navigate Ensembl:
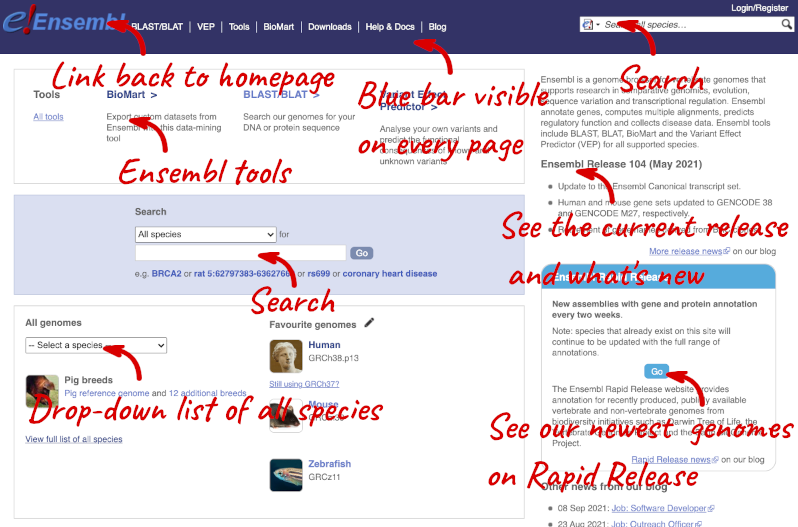
At the top left you can see the current release number and what has come out in this release. To access old releases, scroll to the bottom of the page and click on View in archive site.
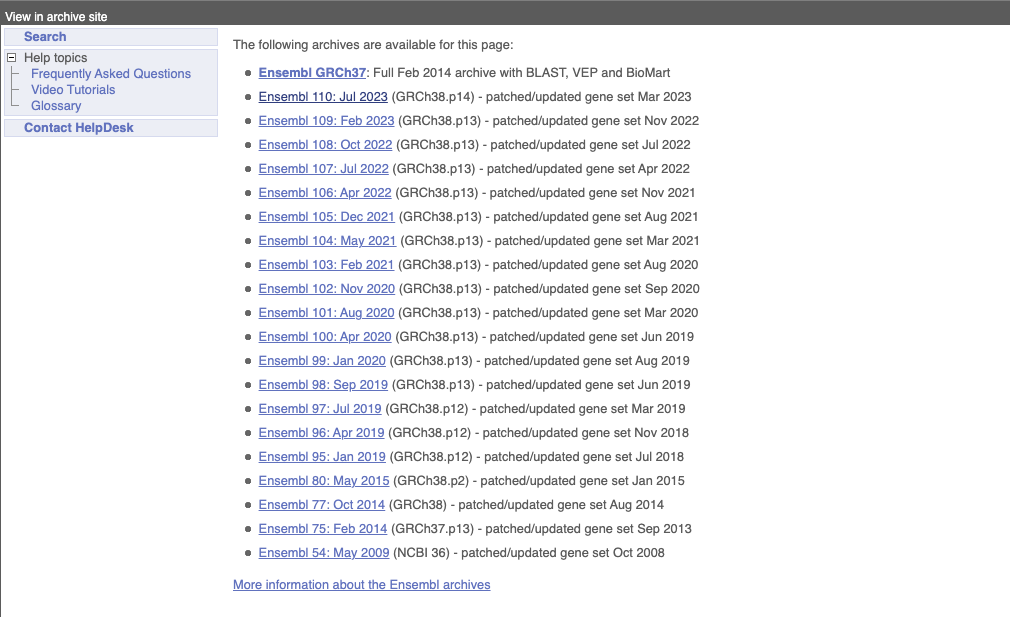
Click on the links to go to the archives. Alternatively, you can jump quickly to the correct release by putting it into the URL, for example e98.ensembl.org jumps to release 98.
Click on View full list of all species.
Click on the common name of your species of interest to go to the species homepage. We’ll click on Chicken.

Here you can see links to example pages and to download flatfiles. To find out more about the genome assembly and genebuild, click on More information and statistics.
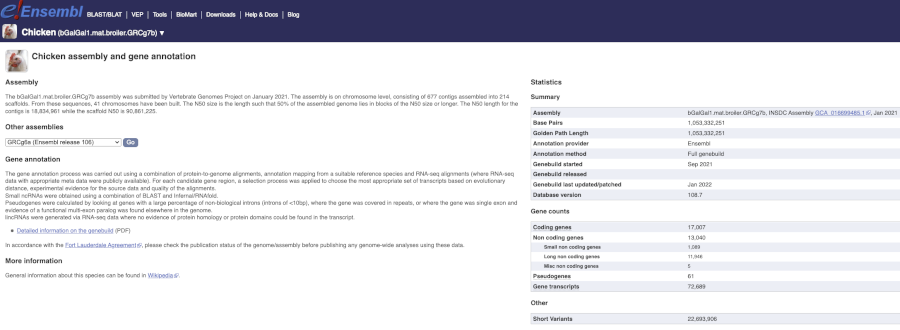
Here you’ll find a detailed description of how to the genome was produced and links to the original source. You will also see details of how the genes were annotated.
Chicken assembly
When was the current Gallus gallus genome assembly submitted and by whom?
Select Chicken from the drop down species list, or click on View full list of all Ensembl species, then choose Chicken from the list to go to the species homepage. Click on on More information and statistics.
The bGalGal1.mat.broiler.GRCg7b assembly was submitted by Vertebrate Genomes Project on January 2021.
Pig species data
-
How many coding and non-coding genes does pig have?
-
When was the current Sus scrofa genome assembly produced and by whom?
1.Select Pig from the drop down species list, or click on View full list of all Ensembl species, then choose Pig from the list to go to the species homepage. Click on More information and statistics.
Pig has 22,063 coding genes and 13,154 non-coding genes.
- The Sscrofa11.1 assembly of the pig genome was produced in January 2017 by the Swine Genome Sequencing Consortium (SGSC).
Exploring genomic regions
Start at the Ensembl front page, ensembl.org. You can search for a region by typing it into a search box, but you have to specify the species.
Type (or copy and paste) chicken 4:53544500-53598000 into either search box.
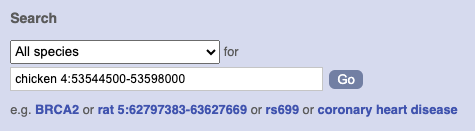
Press Enter or click Go to jump directly to the Region in detail Page.

Click on the  button to view page-specific help. The help pages provide text, labelled images and, in some cases, help videos to describe what you can see on the page and how to interact with it.
button to view page-specific help. The help pages provide text, labelled images and, in some cases, help videos to describe what you can see on the page and how to interact with it.
The Region in detail page is made up of three images, let’s look at each one in detail.
- The first image shows the chromosome:

You can jump to a different region by dragging out a box in this image. Drag out a box on the chromosome, a pop-up menu will appear.

If you wanted to move to the region, you could click on Jump to region (### bp). If you wanted to highlight it, click on Mark region (###bp). For now, we’ll close the pop-up by clicking on the X on the corner.
- The second image shows a 1Mb region around our selected region. This view allows you to scroll back and forth along the chromosome.
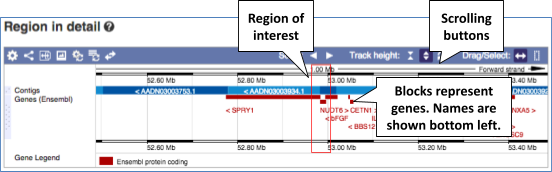
You can also drag out and jump to or mark a region.

Click on the X to close the pop-up menu.
Click on the Drag/Select button  to change the action of your mouse click. Now you can scroll along the chromosome by clicking and dragging within the image. As you do this you’ll see the image below grey out and two blue buttons appear. Clicking on Update this image would jump the lower image to the region central to the scrollable image. We want to go back to where we started, so we’ll click on Reset scrollable image.
to change the action of your mouse click. Now you can scroll along the chromosome by clicking and dragging within the image. As you do this you’ll see the image below grey out and two blue buttons appear. Clicking on Update this image would jump the lower image to the region central to the scrollable image. We want to go back to where we started, so we’ll click on Reset scrollable image.
The third image is a detailed, configurable view of the region.

Click on the Drag/Select option at the top or bottom right to switch mouse action. On Drag, you can click and drag left or right to move along the genome, the page will reload when you drop the mouse button. On Select you can drag out a box to highlight or zoom in on a region of interest.
With the tool set to Select, drag out a box around an exon and choose Mark region.

The highlight will remain in place if you zoom in and out or move around the region. This allows you to keep track of regions or features of interest.
We can edit what we see on this page by clicking on the blue Configure this page menu at the left.

This will open a menu that allows you to change the image.
You can put some tracks on in different styles; more details are in this FAQ: http://www.ensembl.org/Help/Faq?id=335.

Let’s add some tracks to this image. Add:
- All variants on genotyping chips - short variants (SNPs and indels) - Normal
Now click on the tick in the top left hand to save and close the menu. Alternatively, click anywhere outside of the menu. We can now see the tracks in the image.
We can also change the way the tracks appear by hovering over the track name then the cog wheel to open a menu. We can move tracks around by clicking and dragging on the bar to the left of the track name.

Now that you’ve got the view how you want it, you might like to show something you’ve found to a colleague or collaborator. Click on the Share this page button to generate a link. Email the link to someone else, so that they can see the same view as you, including all the tracks you’ve added. These links contain the Ensembl release number, so if a new release or even assembly comes out, your link will just take you to the archive site for the release it was made on.
To return this to the default view, go to Configure this page and select Reset configuration at the bottom of the menu.
Exploring a genomic region in Sus scrofa (Pig)
-
Go to the region from 8,805,953 to 8,858,418 on pig chromosome 11.
-
Configure this page to turn on the Tandem repeats (TRF) track in this view. What is this track? How many TRF overlap this region?
-
Create a URL for this display. Email it to your neighbour.
-
Export the genomic sequence of the region you are looking at in FASTA format.
-
Turn off all tracks you added to the Region in detail page.
-
Go to the Ensembl homepage. Select Pig from the drop-down menu in the blue box and enter
11:8,805,953-8,858,418in the text box. Click Go. - Click Configure this page in the left-hand menu (or on Add/remove tracks at the top left-hand corner of the Region in detail image). Type TRF into the search field in the top left-hand corner of the pop-up menu. Enable the Tandem repeats (TRF) track on the right. You can click on the i icon on the far left for a track description.
The TRF track locates adjacent copies of a pattern of nucleotides. Save and close the new configuration by clicking on the check icon in the top right-hand corner of the pop-up menu or by clicking anywhere outside the pop-up menu. There are 19 TRF that overlap this region.
-
Click Share this page in the left hand-side panel. Copy the URL, get your neighbour’s email address and send them the URL you copied. When you receive the link from them, open the email and click on your link. You should be able to view the page with the new configuration and data tracks they have added.
-
Click on Export data in the left-hand menu. Leave the default parameters as they are. Click Next> and view the sequence in a new browser tab by clicking on Text. The sequence is in FASTA formatwhich comprises a header (beginning with >) that provides information about the genome assembly (primary_assembly:Sscrofa11.1), the chromosome, the start and end coordinates and the strand. For example:
>primary_assembly:Sscrofa11.1:11:8805953-8858418:1 - Click on Reset configuration at the top of the Region in detail image.
Exploring a genomic region in Gallus gallus (Chicken)
-
Go to the region from 38,111,022-38,265,293 on chicken chromosome 5. How many contigs make up this portion of the assembly (contigs are contiguous stretches of DNA sequence that have been assembled solely based on direct sequencing information)?
-
Zoom in on ESRRB gene.
-
Turn on the RefSeq GFF3 annotation track as Expanded with labels.
-
Save this image in PDF format.
- Go to the Ensembl homepage. Select Chicken from the drop-down menu in the blue box and enter 5:38111022-38265293 into the text box. Click Go.
This genomic region is made up of one contig indicated by the dark blue coloured bar in the Contigs track.
-
Make sure your cursor is set to the Select a region action (you can change your cursor action in the top right-hand corner of the Region in detail view). Drag a box around the ESRRB gene (note that you will need to highlight the feature itself, i.e. the block, rather than the label) and click on Jump to region.
-
Click on Configure this page in the left-hand panel to open the configuration menu. Enter RefSeq GFF3 annotation into the search box in the top left-hand corner. To enable the track, click on the square next to the track name RefSeq GFF3 annotation and select the Expanded with labels style. Save and close the pop-up menu.
- Click on the Export this image icon above the image and then on the Download button to download the image in PDF format.
Genes and Transcripts
Exploring the MYH9 gene in Gallus gallus (Chicken)
- Find the MYH9 (myosin, heavy chain 9, non-muscle) gene in the chicken reference, and go to the Gene tab.
- On which chromosome and which strand of the genome is this gene located?
- Which transcript produces the longest protein and how long is the protein sequence?
-
What are some functions of MYH9 according to the Gene Ontology consortium? Have a look at the GO pages for this gene.
- In the transcript table, click on the transcript ID for MYH9-209, and go to the Transcript tab.
- How many exons does it have?
- Are any of the exons completely or partially untranslated?
- Is there an associated sequence in UniProt? Have a look at the General identifiers for this transcript.
- Are there microarray (oligo) probes that can be used to monitor ENSGALT00010036169.1 expression?
- Go to the Ensembl homepage. Select Chicken from the drop-down list in the blue box, enter MYH9 and click Go. In the search results page, click on Chicken reference in the left-hand panel to restrict your results to the reference genome only. Click on the first hit MYH9 (Chicken Gene, Breed: reference) to open the Gene tab. Look at the Location section in the gene summary at the top of the page.
The gene is located on chromosome 1 on the forward strand.
Now click on the Show transcript table button and focus on the Protein column in the Transcript table.
The transcript ENSGALT00010036169.1 (MYH9-209) produces the longest protein at 1,960 amino acid residues.
- Gene Ontology maps terms to a protein in three classes: biological process, cellular component, and molecular function.
Meiotic spindle organisation, cell morphogenesis, and angiogenesis are some of the roles associated with the MYH9 gene.
- Click on ENSGALT00010036169.1 in the Transcript table to open the corresponding Transcript tab. Look at the About this transcript section in the transcript summary at the top of the page.
The transcript has 41 exons.
Click on the Exons link in the left-hand side menu. In the Sequence column of the Exon table, look for any UnTranslated Regions (UTRs) which coloured in orange.
Exon 1 is completely untranslated, and exons 2 and 41 are partially untranslated. You can also see this in the cDNA view if you click on Sequence: cDNA in the left-hand menu.
Click on External References: General identifiers in the left-hand menu. Look for UniProtKB in the External database column.
A0A1D5PM19.34 from UniProt matches the translation of the Ensembl transcript. Click on A0A1D5PM19.34 to open the corresponding UniProt entry in a new browser tab.
- In the left-hand menu, look for External References: Oligo probes.
There are probes from Affy and Agilent that can be used to monitor expression of this transcript.
Comparative genomics
Let’s look at the homologues of the pig BRCA2 gene. Search for the gene and go to the Gene tab.
Click on Gene tree to display the current gene in the context of a phylogenetic tree used to determine orthologues and paralogues.
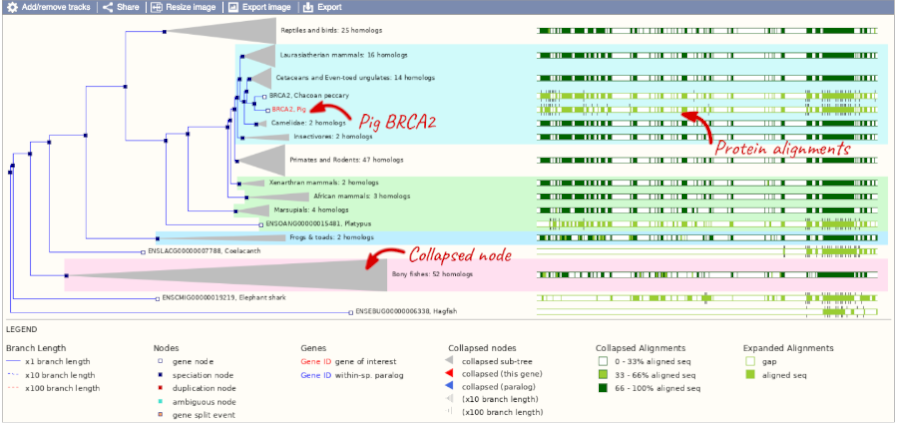
You can change the gene tree display by using the View options below the image, the Configure this page menu as well as menus for individual nodes, which you can open by clicking on the nodes. Grey funnels indicate collapsed nodes. You can expand them by clicking on the node and selecting expand this sub-tree from the pop-up menu.
You can download the tree in a variety of formats. Click on the download icon in the bar at the top of the image to get a pop-up window where you can choose your format.

You can look at homologues in the Orthologues and Paralogues pages, which can be accessed from the left-hand menu. If there are no orthologues or paralogues, then the option will be greyed out. Paralogues is greyed out for BRCA2 indicating that there are no paralogues.
Click on Orthologues to see the available orthologues.
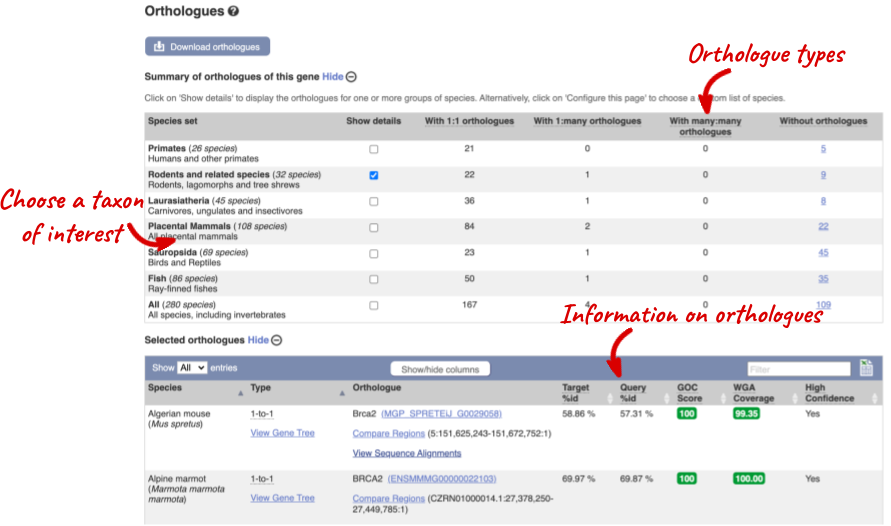
Choose to see only Rodent and related species orthologues by selecting the box. The table below now only shows details of these orthologues. Let’s look at mouse (Mus musculus).
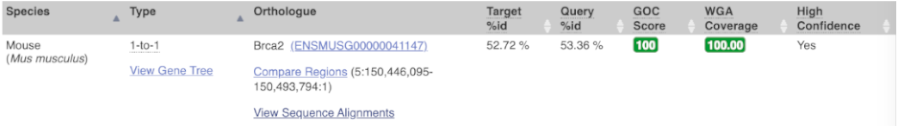
Links from the orthologue allow you to go to alignments of the orthologous proteins and cDNAs. Click on View Sequence Alignments, and then View Protein Alignment in the pop-up menu for the mouse orthologue.
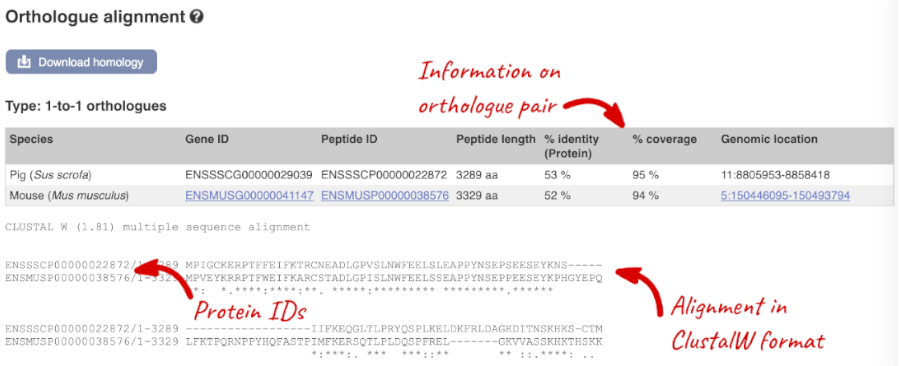
Let’s look at some of the comparative genomics views in the Location tab. Go to the region 15:81873000-82000800 in pig, which contains the HoxD cluster which is involved in limb development and is highly conserved between species.
You can turn on conservation scores and constrained elements. Click on Configure this page, then Comparative genomics and turn on the tracks for Constrained elements for 16 pig breeds EPO-Extended and Conservation score for 16 pig breeds EPO-Extended. Save and close the menu.
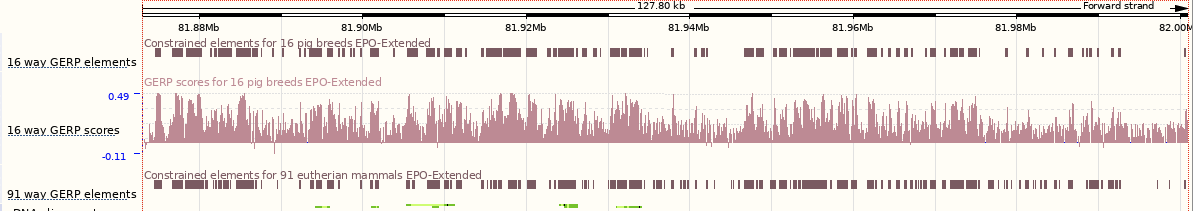
You can now see the conservation scores in pale pink. These were used to determine the peaks indicated in the constrained elements track in dark pink. This track indicates regions of high conservation between species, considered to be “constrained” by evolution.
We can also look at individual species comparative genomics tracks in this view by clicking on Configure this page.
Select BLASTz/LASTz alignments from the left-hand menu to choose alignments between closely related species. Turn on the alignments for_Human_ and Cow in Normal. Save and close the menu.

We can also look at the alignment between species or groups of species as text. Click on Alignments (text) in the left hand menu.
Select Cow from the alignments list then click Go.
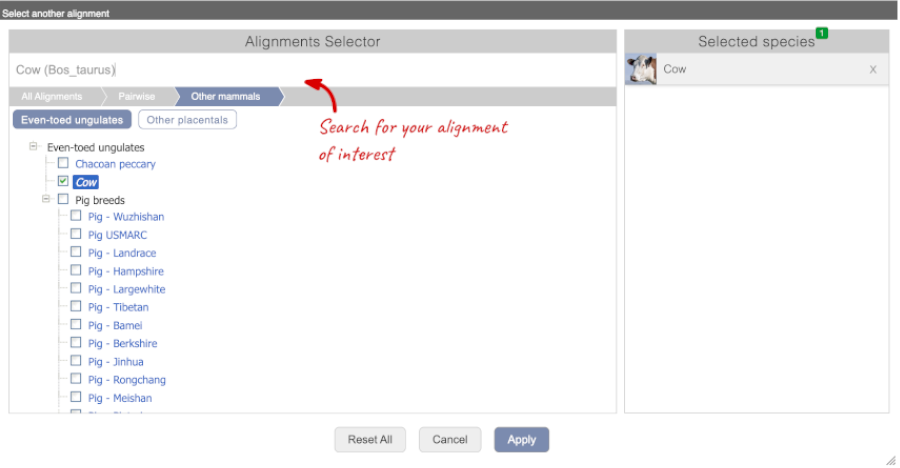
You will see a list of the regions aligned, followed by the sequence alignment. Exons are shown in red.
To compare with both contigs visually, go to Region comparison.
To add species to this view, click on the blue Select species or regions button. Choose Cow from the list then close the menu.
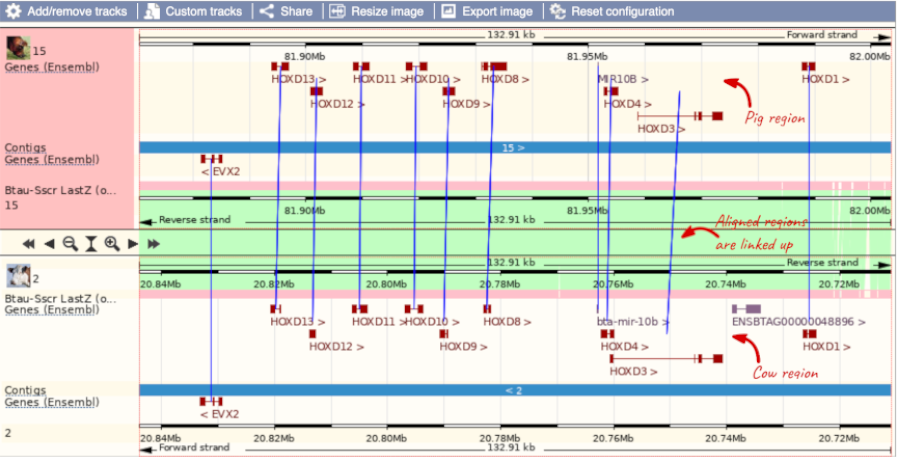
You can configure this view for both species. Click on configure this page and look in the top left of the menu.
The drop down allows you to configure each species separately.
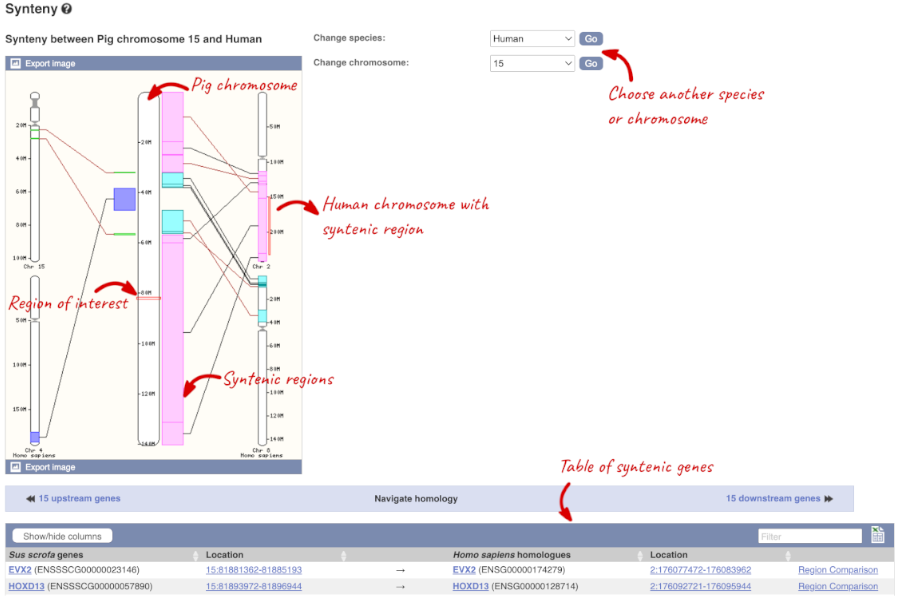
We can view large scale syntenic regions from our chromosome of interest. Click on Synteny in the left hand menu.
Orthologues and gene trees for the Gallus gallus (Chicken) BRAF gene
- Let’s explore the orthologues of the chicken BRAF gene.
- How many orthologues are predicted for the chicken BRAF in sauropsida (birds and reptiles)?
- How much sequence identity does the Anolis carolinensis (Green anole) protein have to the chicken one?
- Export the protein alignment in Clustal format.
- Look at the orthologue in human. Is there a genomic alignment between human and chicken? Is there a gene for both species in this region?
- Go to the Ensembl homepage, select Chicken from the drop-down list in the blue search box, enter BRAF and click Go. Open the **Gene tab and click on Comparative Genomics: Orthologues at the left-hand panel to see all the orthologous genes.
There 25x 1:1 and 1x 1:many orthologues in sauropsida.
Find Green anole in the Selected orthologues (you can use the filter in the top right-hand corner).
The percentage of identical amino acids in the Green anole protein (the orthologue) compared with the gene of interest. i.e. chicken BRAF (the target species/gene) is 84.42%. This is known as the Target %id. The identity of the gene of interest (chicken BRAF) when compared with the orthologue (Green anole BRAF, the query species/gene) is 94.67% (this is the Query %id).
Click on the View Sequence Alignments link in the Orthologue column of the Selected orthologues table and select View Protein Alignment in the pop-up menu. To download the alignment, click on the Download homology button and select the CLUSTALW file format in the pop-up menu.
- Click on Comparative Genomics: Genomic alignments in the left-hand panel. Click on Select an alignment and add Human in the pop-up menu. In the table, select Block 1 to view the largest block of aligned sequence (this will lead you to the Location tab). Click on Display full alignment. In the alignment, sequences coloured in red are exons.
There is a gene in both species in this region. You can find where the start and stop codons are located if you Configure this page and select Codons: START/STOP codons in the options.
Note: You can visualise the alignment in the genomic context in the Comparative Genomics: Region Comparison page (blue lines connect homologous genes between species). Go to Select species or regions, add Human and close the pop-up menu. Click on Configure this page. In the pop-up menu under Comparative features category, enable the Join genes option. You may need to zoom out on the Region in detail view to see blue lines connecting all the homologous genes between chicken and human genes in that region.
Whole-genome alignments in Sus scrofa (Pig)
Go to www.ensembl.org to find the DBH gene on the reference pig genome (Sscrofa11.1).
-
Go to the Location page for this gene. View the Alignments (image) and Alignments (text) for the 16 pig breeds EPO-Extended. Do all the pig breeds show a gene in these alignments?
-
Export the alignments in ClustalW format.
-
Go to the Region in detail view and turn on the 16 pig breeds EPO-Extended multiple alignment, conservation score and constrained elements tracks. Are there any differences between the conservation score and constrained elements tracks?
-
Compare the 16 way GERP elements track and the 91 way GERP elements track that is already turned on by default.
- What is the difference between the two tracks?
- Which regions of the gene do most of the constrained element blocks match-up to?
- How can you find more information on how the constrained elements track was generated?
- Search for the DBH(ENSSSCG00000005742) gene in the Pig (SScrofa11.1) reference and switch to the Location tab. Click on Alignments (image) in the left-hand panel. Under Alignment, click on the Select alignment button to open a pop-up menu. Enable the 16 pig breeds EPO-Extended alignment, then close the menu.
All 12 big breeds as well as cow, horse and sheep have an alignment at this region. This can also be seen in the Alignments (text) page, where the exons are coloured in red.
-
You can export the alignments from either the Alignments (images) or Alignments (text) pages. Click on the blue Download alignments button at the top of the page. From the pop-up menu, select File format: CLUSTALW. You can Preview the alignment in a new browser tab, or Download the file to your local machine.
- Click on Region in detail in the left-hand panel. In the pop-up menu, go to the Comparative genomics section and turn on the following tracks:
- Multiple alignments: 16 pig breeds EPO-Extended
- Conservation score for 16 pig breeds EPO-Extended
- Constrained elements for 16 pig breeds EPO-Extended
Close the pop-up menu and find the tracks in the Region in detail view.
The 16 pig breeds EPO-extended track shows that the entire region for the DBH gene can be aligned among the big breeds and related agricultural species. The Constrained elements and Conservation score tracks show where the conserved sequence is located in the alignment. Regions where constrained elements are found are regions with high GERP scores. Higher conservation regions (i.e. constrained elements) match up with exonic regions (exons tend to be highly conserved) of the gene. Note that there are intronic regions that seem to be fairly conserved across the species available.
- For both 16 way GERP elements and 91 way GERP elements tracks, click on the track name to open the pop-up menu. Hover over the i icon with your cursor to find a track description.
The 16 way GERP elements track shows the 16 pig breeds EPO-Extended multiple whole-genome alignment. The 91 way GERP elements track shows the 91 eutherian mammals EPO-Extended multiple whole-genome alignment.
You can move the 91 way GERP elements track closer to the 16 way GERP elements track to make any comparisons easier.
You will notice that constrained elements match-up with exonic regions in the genome.
Click on the track name and open the information tab in the pop-up menu. Click on the GERP conservation score link.
This opens the documentation page for the multiple whole-genome alignment calculations.
BioMart
Follow these instructions to guide you through BioMart to answer the following query:
You have three questions about a set of chicken genes:
ESPN, MYH9, USH1C, CISD2, THRB, WHRN
(these are HGNC gene symbols. More details on the HUGO Gene Nomenclature Committee can be found on https://www.genenames.org/)
- What are the NCBI Gene IDs for these genes?
- Are there associated functions from the GO (gene ontology) project that might help describe their function?
- What are their cDNA sequences?
Click on BioMart in the top header of the Ensembl website or go to BioMart directly by visiting https://www.ensembl.org/biomart/martview.
You cannot choose any filters or attributes until you’ve chosen your dataset. Your dataset is the data type you’re working with. In this case we’re going to choose genes, so pick Ensembl Genes then Chicken genes from the drop-downs.
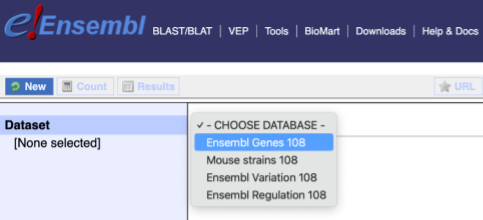
Now that you’ve chosen your dataset, the filters and attributes will appear in the column on the left. You can pick these in any order and the options you pick will appear.
Click on Filters on the left to see the available filters appear on the main page. You’ll see that there are loads of categories of Filters to choose from. You can expand these by clicking on them. For our query, we’re going to expand GENE.
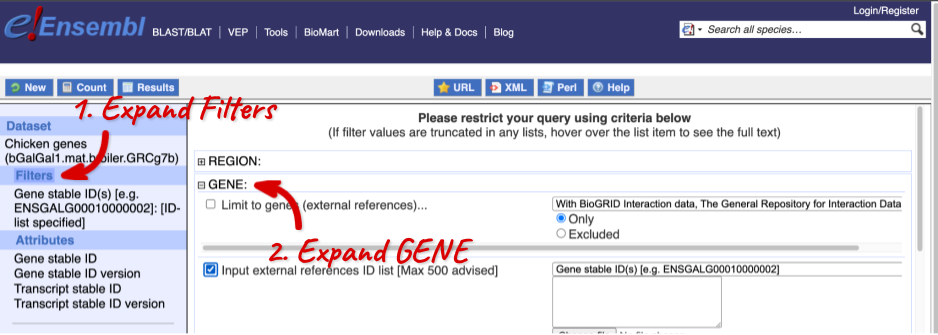
Our input data is a list of identifiers, so we’re going to use the Input external references ID list filter. This allows us to input a list of identifiers from different databases. We need to choose what kind of identifier we’re using, so that BioMart can look up the right column in a data table. You can pick these from a drop-down list, which lists the type of identifier with an example of how it looks. For our query, we have a list of gene names, so we need to pick Gene Name(s).

To check if the filters have worked, you can use the Count button at the top left, which will show you how many genes have passed the filter. If you get 0 or another number you don’t expect, this can help you to see if your query was effective.
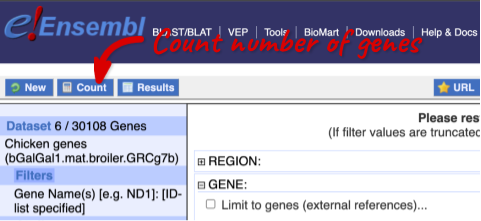
To choose the attributes, expand this in the menu. There are six categories for chicken gene attributes. These categories are mutually exclusive, you cannot pick attributes from multiple categories. This means that we need to do two separate queries to get our GO terms and NCBI IDs, and to get our cDNA sequences.
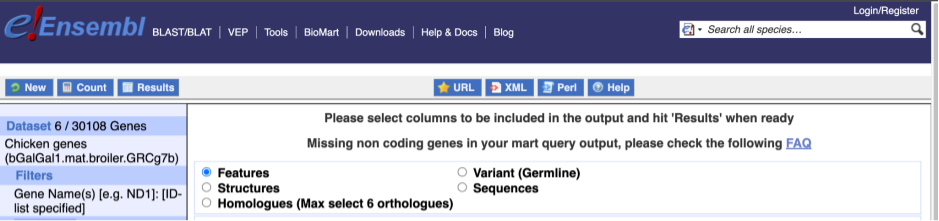
The Ensembl gene and transcript IDs, with and without version numbers are selected by default. The selected attributes are also listed on the left.
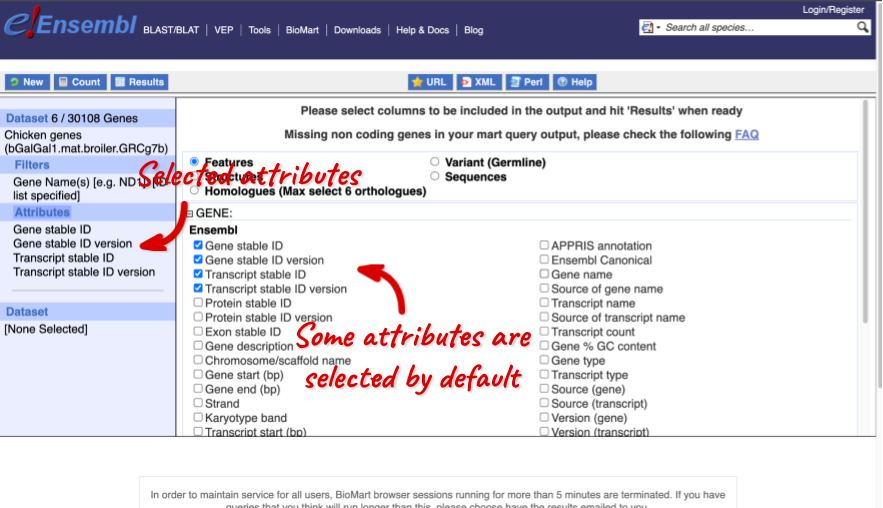
We can choose the attributes we want by clicking on them. For our query, we’re going to select:
- GENE
- Gene Name
- EXTERNAL
- NCBI gene ID
- GO term accession
- GO term name
- GO term definition
We need to select the Gene Name in order to get back our original input, as this is not returned by default in BioMart. The order that you select the attributes in will define the order that the columns appear in in your output table.
You can get your results by clicking on Results at the top left.
The results table just gives you a preview of the first ten lines of your query. This allows the results to load quickly, so that if you need to make any changes to your query, you don’t waste any time. To see the full table you can click on View ## rows. You can also export the data to an xls, tsv, csv or html file. For large queries, it is recommended that you export your data as Compressed web file (notify by email), to ensure your download is not disrupted by connection issues.
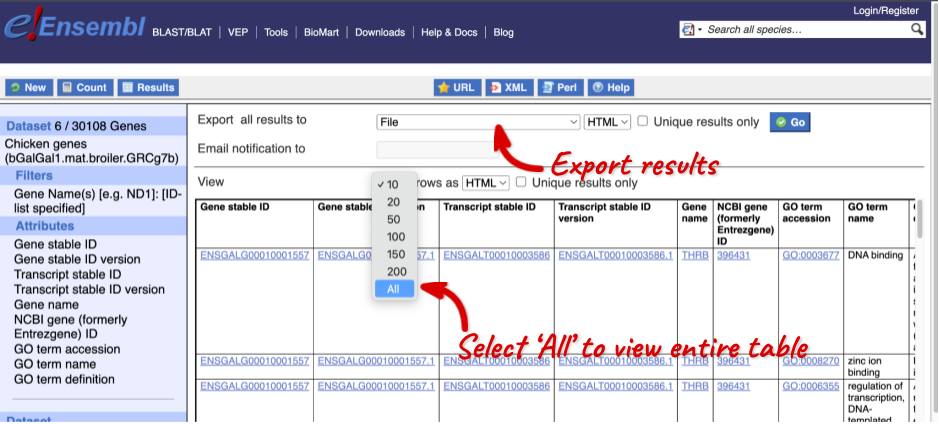
You can see multiple rows per gene in your input list, because there are multiple transcripts per gene and multiple GO terms per transcript.
To get the cDNA sequences, go back to the Attributes then select the category Sequences and expand SEQUENCES.
When you select the sequence type, the part of the transcript model you’ve chosen will be highlighted in the grpahic.
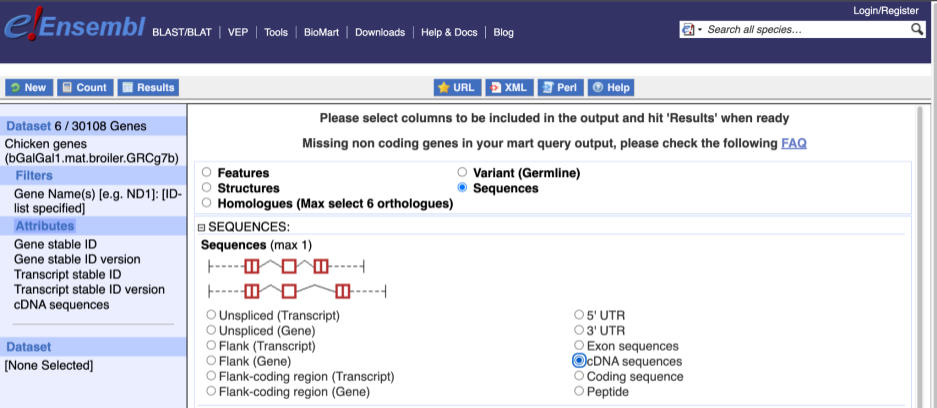
Choose cDNA sequences, then expand HEADER INFORMATION to add Gene Name to the header. Then hit Results again.
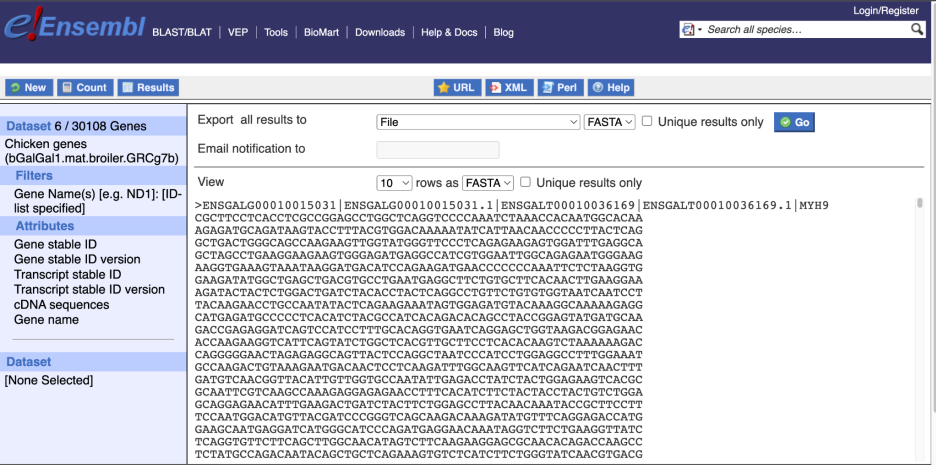
For more details on BioMart, have a look at this publication: Kinsella RJ, Kähäri A, Haider S, et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database: the Journal of Biological Databases and Curation. 2011; 2011:bar030. DOI: 10.1093/database/bar030. PMID: 21785142; PMCID: PMC3170168.
BioMart: Convert IDs
BioMart is a very handy tool when you want to convert IDs from different databases. The following is a list of 27 IDs of Sus scrofa proteins from the NCBI RefSeq database: NP_001116455,NP_001231885,NP_001230616,NP_001231413,NP_001231746,NP_999129,NP_001231602,NP_001177096,NP_001231419,NP_001230512, NP_001231165,NP_001167636,NP_001172069,NP_001011509,NP_999191,NP_001231786,NP_001231468,NP_001121951,NP_001230557,NP_999413
Generate a list that shows to which Ensembl Gene IDs and to which gene names these RefSeq IDs correspond. Do these 27 proteins correspond to 27 genes?
Click New. Choose the ENSEMBL Genes database. Choose the Pig genes (Sscrofa11.1) dataset.
Click on Filters in the left panel. Expand the GENE section by clicking on the + box. Select Input external references ID list - RefSeq peptide ID(s) and enter the list of IDs in the text box (either comma separated or as a list). HINT: You may have to scroll down the menu to see these. Count shows 20 genes.
Click on Attributes in the left panel. Select the Features attributes page. Expand the GENE tab by clicking on the + box. Select Gene name. Expand the EXTERNAL tab. Select RefSeq Peptide ID.
Click the Results button on the toolbar. Select View All rows as HTML or export all results to a file.
BioMart: Finding genes by protein domain
Find chicken proteins with transmembrane domains located on chromosome 9.
As with all BioMart queries you must select the dataset, set your filters (input) and define your attributes (desired output). For this exercise:
Dataset: Ensembl genes in chicken
Filters: Transmembrane proteins on chromosome 9
Attributes: Ensembl gene and transcript IDs and Associated gene names
Go to the Ensembl homepage (https://www.ensembl.org) and click on BioMart at the top of the page. Select Ensembl genes as your database and Chicken genes (bGalGal1.mat.broiler.GRCg7b) as the dataset. Click on Filters on the left of the screen and expand REGION. Change the chromosome to 9. Now expand PROTEIN DOMAINS AND FAMILIES, also under filters, and select Limit to genes …, choosing With Transmembrane helices from the drop-down and select Only. Clicking on Count should reveal that you have filtered the dataset down to 143 genes.
Click on Attributes. Under Features expand GENE. Select Gene name.
Now click on Results. The first 10 results are displayed by default; display all results by selecting All from the drop-down menu above the table.
The output will display the Ensembl gene ID, Ensembl Transcript ID and associated gene names of all proteins with a transmembrane domain on chicken chromosome 9. If you prefer, you can also export as an Excel sheet by using the Export all results to XLS option.
BioMart: Find genes associated with array probes
Here are two affymetrix probeset IDs from my microarray experiment that seem to map uniquely to genes in the chicken genome: Gga.12669.1.S1_at, GgaAffx.7784.1.S1_at
(a) Retrieve for the genes corresponding to these probe-sets the Ensembl Gene and Transcript IDs as well as their gene symbols and descriptions.
(b) In order to analyse these genes for possible promoter/enhancer elements, retrieve the 2000 bp upstream of the transcripts of these genes.
(c) In order to be able to study these chicken genes in duck, identify their duck orthologues. Also retrieve the genomic coordinates of these orthologues.
(a) Click New. Choose the Ensembl Genes database. Choose the Chicken genes dataset.
Click on Filters in the left panel. Expand the GENE section by clicking on the + box. Select Input microarray probes/probesets ID list - AFFY Chicken probe ID(s) and enter the list of probeset IDs in the text box (either comma separated or as a list).
Count shows three genes match this list of probesets.
Click on Attributes in the left panel. Select the Features attributes page. Expand the GENE section by clicking on the + box. In addition to the default selected attributes, select Gene name and Gene description. Expand the EXTERNAL section by clicking on the + box. Select AFFY Chicken probe from the Microarray probes/probesets section.
Click the Results button on the toolbar. Select View All rows as HTML or export all results to a file. Tick the box Unique results only.
Your results should show that the 2 probes map to 2 Ensembl genes.
(b) Don’t change Dataset and Filters – simply click on Attributes.
Select the Sequences category. Expand the SEQUENCES tab by clicking on the + box. Select Flank (Transcript) and enter 2000 in the Upstream flank text box. Expand the HEADER INFORMATION tab by clicking on the + box. Select Gene description and Gene name in addition to the default selected attributes.
Note: Flank (Transcript) will give the flanks for all transcripts of a gene with multiple transcripts. Flank (Gene) will give the flanks for one possible transcript in a gene (the most 5’ coordinates for upstream flanking).
Click the Results button on the toolbar.
(c) You can leave the Dataset and Filters the same, and go directly to the Attributes section:
Click on Attributes in the left panel. Select the Homologues category. Expand the GENE tab by clicking on the + box. Select Gene name. Unselect Transcript stable ID and Transcript stable ID version. Expand the ORTHOLOGUES [A-E] tab by clicking on the + box. Select Duck gene stable ID, Duck chromosomes/scaffold name, Duck chromosome/scaffold start (bp) and Duck chromosome/scaffold end (bp).
Click the Results button on the toolbar. Select View All rows as HTML or export all results to a file.
Your results should show that for each chicken gene, one duck orthologue has been identified.
Variation
In any of the sequence views shown in the Gene and Transcript tabs, you can view variants on the sequence. You can do this by clicking on Configure this page from any of these views.
Let’s take a look at the Gene sequence view for MCM6 in chicken. Search for MCM6 and go to the Sequence view.
If you can’t see variants marked on this view, click on Configure this page and select Show variants: Yes and show links.
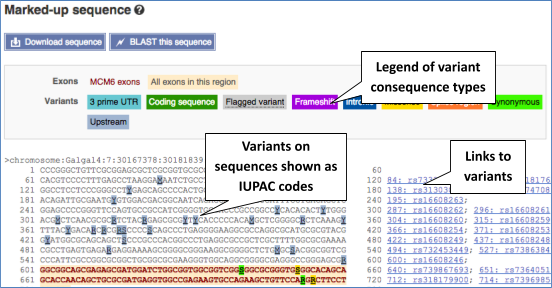
Find out more about a variant by clicking on it.
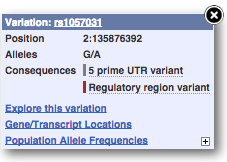
You can add variants to all other sequence views in the same way.
You can go to the Variation tab by clicking on the variant ID. For now, we’ll explore more ways of finding variants.
To view all the sequence variations in table form, click the Variant table link at the left of the gene tab.
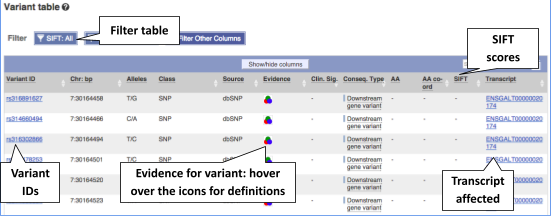
You can filter the table to only show the variants you’re interested in. For example, click on Consequences: All, then select the variant consequences you’re interested in.
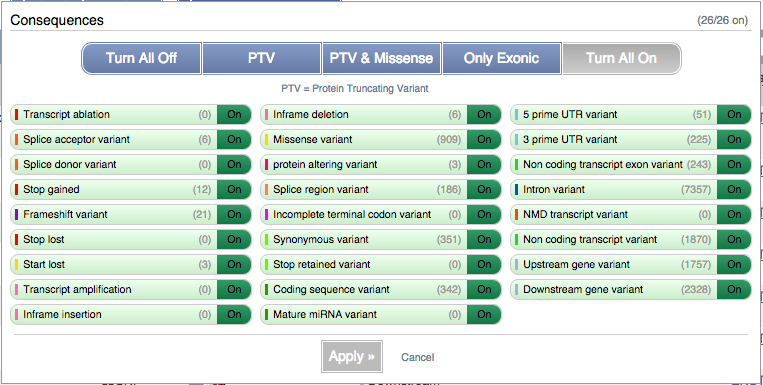
You can also filter by SIFT, or click on Filter other columns for filtering by other columns such as Evidence or Class.
The table contains lots of information about the variants. You can click on the IDs here to go to the Variation tab too.
You can also see the phenotypes associated with a gene. Click on Phenotype in the left hand menu.
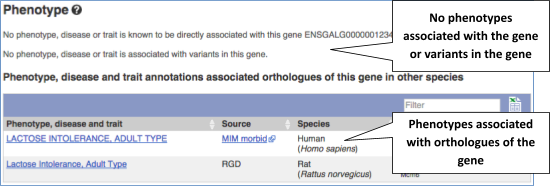
Let’s have a look at variants in the Location tab. Click on the Location tab in the top bar.
Click on Configure this page and open Variation from the left-hand menu.
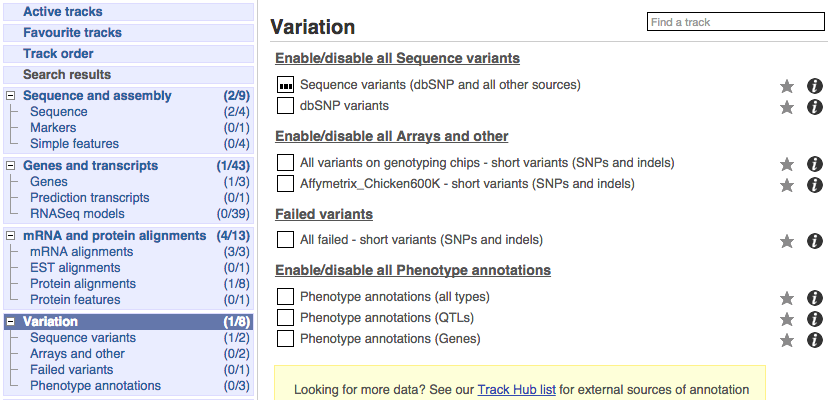
There are various options for turning on variants. You can turn on variants by source, by frequency, presence of a phenotype or by individual genome they were isolated from. You can also turn on QTLs, which cover a locus without being associated with a specific variant. Turn on the following variation tracks.
- All variants on genotyping chips - short variants (SNPs and indels)
- Phenotype annotations (QTLs)

Click on a variant to find out more information. It may be easier to see the individual variants if you zoom in.
Let’s have a look at a specific variant, which happens to fall within the MCM6 gene: rs14625781.
The easiest way to find this variant is if we put rs14625781 into the search box. Click through to open the Variation tab.
The icons show you what information is available for this variant. Click on Genes and regulation, or follow the link at the left.
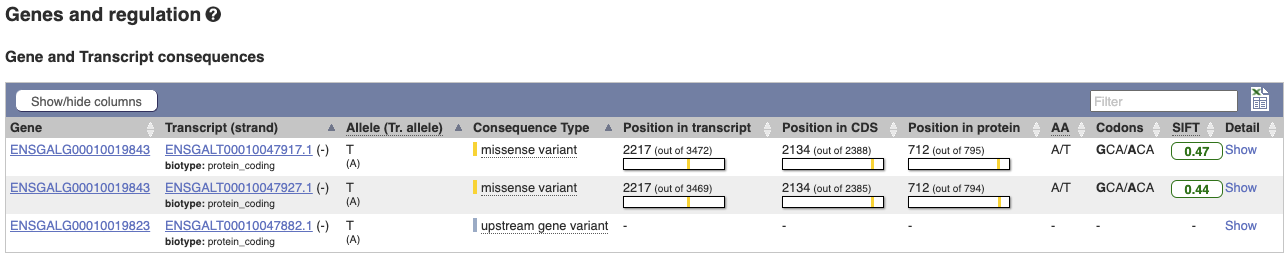
This variant is found in three transcripts of the MCM6 gene, and is missense in two. SIFT predicts that it is unlikely to affect protein function of either (Tolerated).
Let’s look at population genetics. Either click on Explore this variant in the left hand menu then click on the Population genetics icon, or click on Population genetics in the left-hand menu.
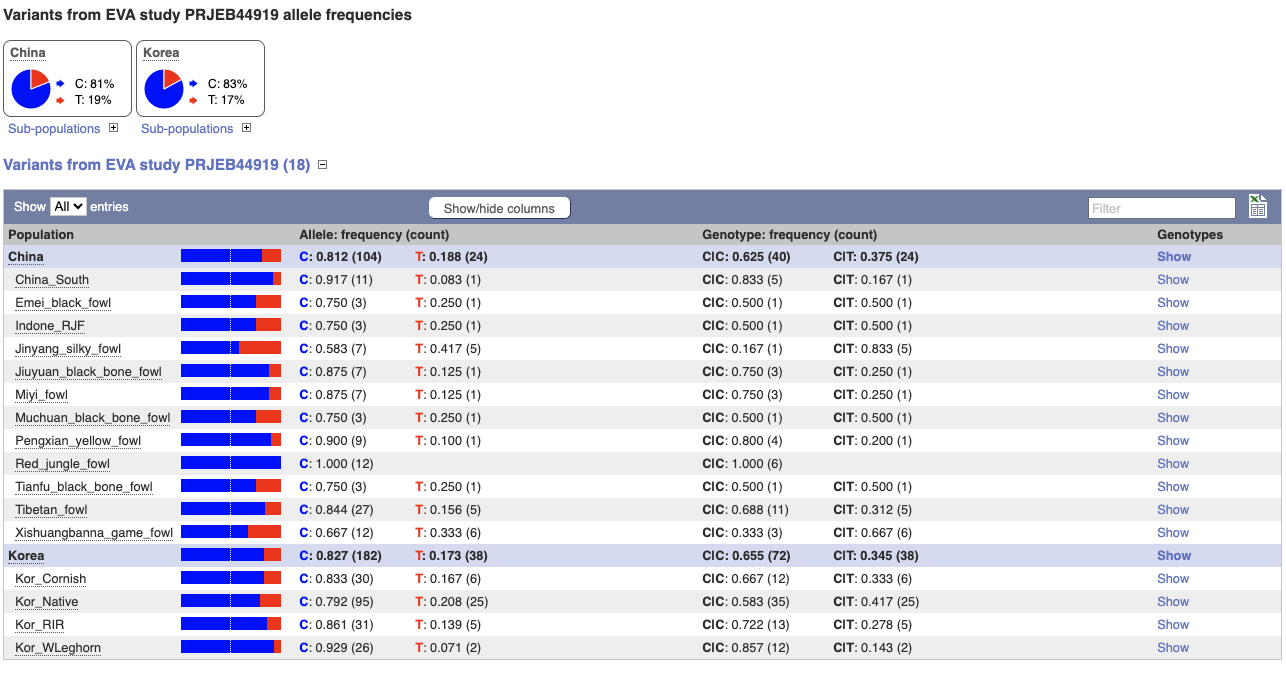
We can see data from EVA study PRJEB44919 showing the frequency of the alleles and genotypes. We can see what animals these genotypes were actually observed in by going to Sample genotypes.
Click on Phylogenetic context to see the variant in other species.
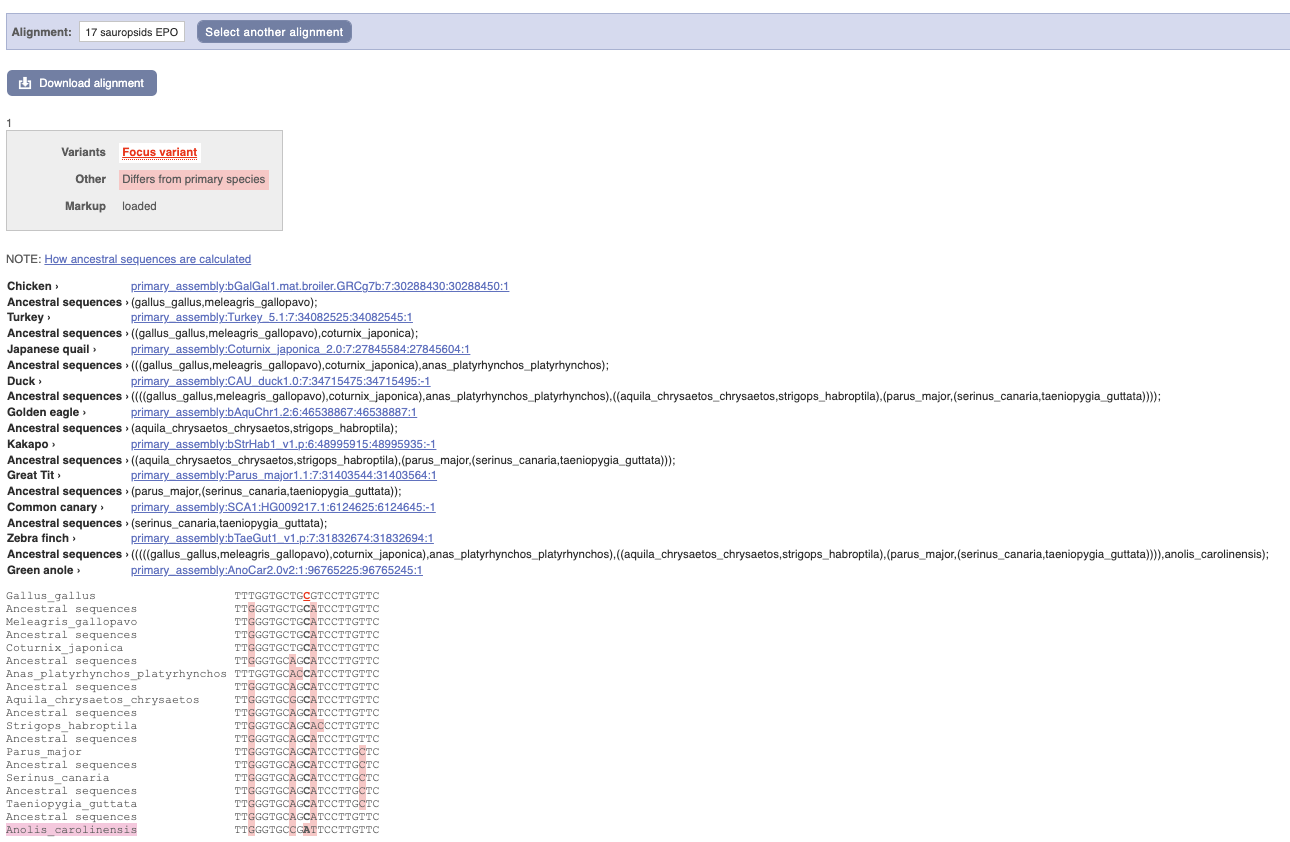
We can see that other birds also have the C alleles as a reference whereas Anolis_carolinensis has an A allele.
Exploring a SNP in chicken
(a) Find the page with information for the chicken SNP rs10731268.
(b) What gene(s) does rs10731268 fall within? What is its effect?
(c) Have any papers been written mentioning rs10731268? What are they about?
(d) What allele is at this position in other birds? What is the likely ancestral allele?
(a) Go to the Ensembl homepage.
Type rs10731268 in the Search box, then click Go. Click on rs10731268.
(b) Click on Genes and Regulation in the side menu (or the Genes and Regulation icon).
rs10731268 falls within 2 genes: ENSGALG00010028562 and ENSGALG00010028568 (HGNC: MLLT1). This variant has a missense consequence in seven transcripts of the ENSGALG00010028562 gene, and downstream gene variant consequence in three transcripts of the ENSGALG00010028568 (HGNC: MLLT1) gene.
(c) Click on Citations in the left hand side menu.
This variant is mentioned in the paper ‘Identification and characterization of genes that control fat deposition in chickens’ from 2013 by D’Andre et al. Click on the PubMed ID 24206759 to go to the paper.
(d) Click on Phylogenetic Context in the side menu. Select Alignment: 17 sauropsids EPO and click Go.
Japanese quail, Duck, Golden Eagle, Common canary and Zebra finch all have an A in this position. This suggests that A may be the ancestral allele.
Exploring a variant in pig
The human gene MC4R has been associated with obesity. The SNP rs81219178 has been identified as a variant in the pig MC4R gene.
(a) What is the amino acid change caused by rs81219178 in MC4R of the pig? Is the change likely to alter the protein function?
(b) How many transcripts does this variant affect? What are the consequences of this variant?
(a) Go to the Ensembl homepage.
Type rs81219178 in the Search box, then click Go.
Click on rs81219178 (Pig Variant, Breed: reference).
Click on Genes and regulation in the left-hand menu or on the icon.
The variant causes a D->N amino acid change (Aspartic acid -> Asparagine). The SIFT score of 0.01 predicts that this change will have a deleterious effect on the protein.
(b) This variant affects one transcript (ENSSSCT00000091644.1) of ENSSSCG00000051798 gene and it has the missense consequence.
Ensembl VEP
We have identified seven variants in pig:
rs319195925, rs80805426, rs81267388, rs80854621, rs711163915, rs321793337, rs792403417
We will use the Ensembl VEP to determine:
- If the variants have been annotated in Ensembl already
- If genes are affected by the variants
Go to the front page of Ensembl and click on Variant Effect Predictor in the Tools section or click on VEP in the top header.

This page contains information about the VEP, including a link for downloading the script version of the tool. Click on the Launch VEP button to open the input form.
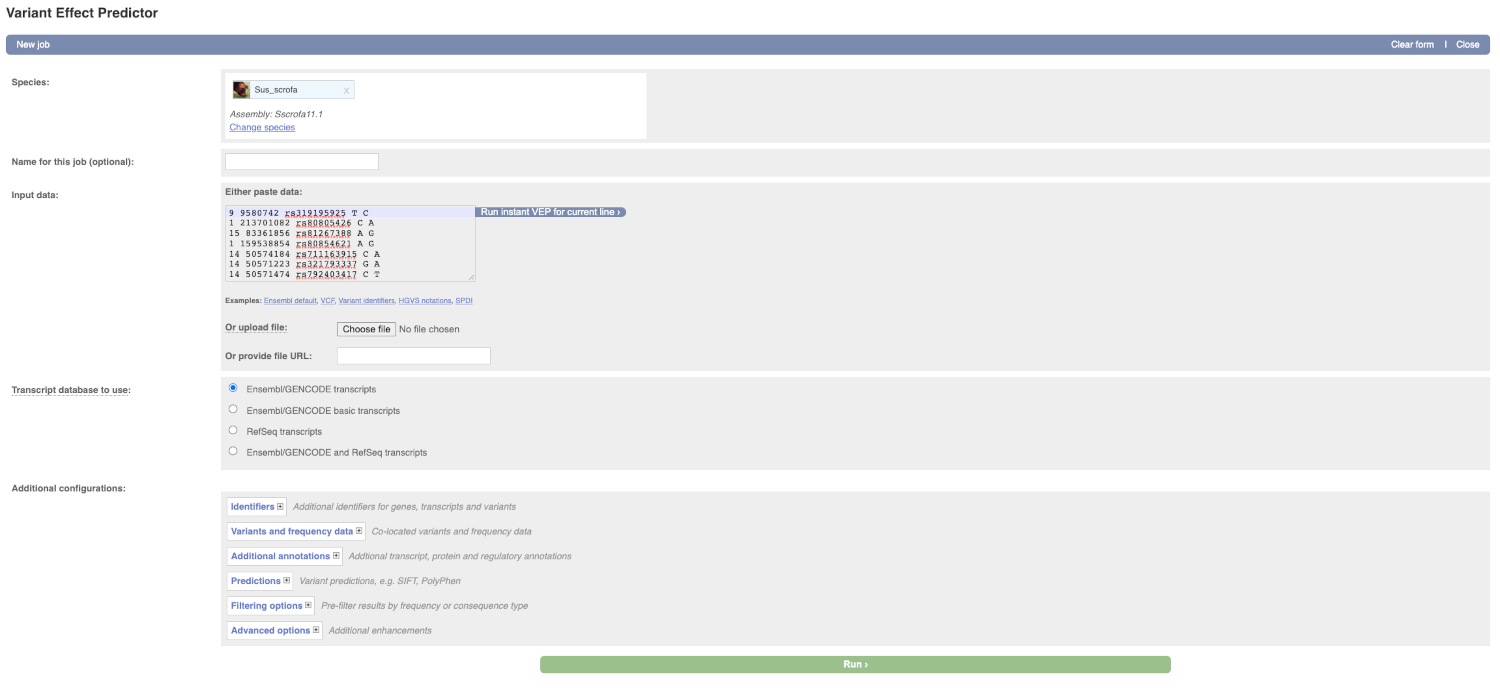
Lets input the variants data in VCF format:
Chromosome Position Name Reference Alternative
Put the following into the Input data box:
9 9580742 rs319195925 T C
1 213701082 rs80805426 C A
15 83361856 rs81267388 A G
1 159538854 rs80854621 A G
14 50574184 rs711163915 C A
14 50571223 rs321793337 G A
14 50571474 rs792403417 C T
The VEP will detect automatically that the data is in VCF format.
There are further options that you can choose for your output. These are categorised as Identifiers, Variants and frequency data, Additional annotation, Predictions, Filtering options and Advanced options. Let’s open all menus and take a look.
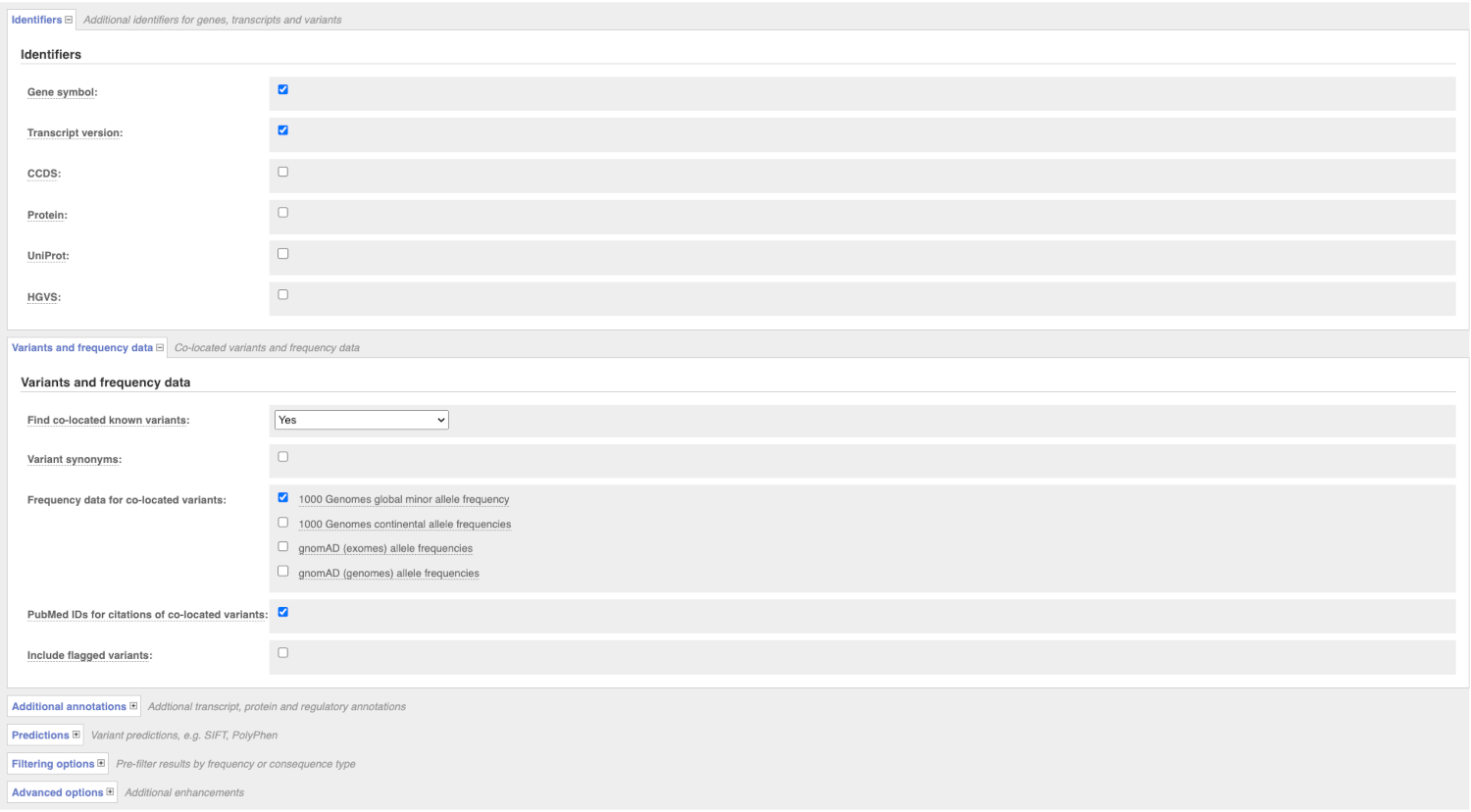
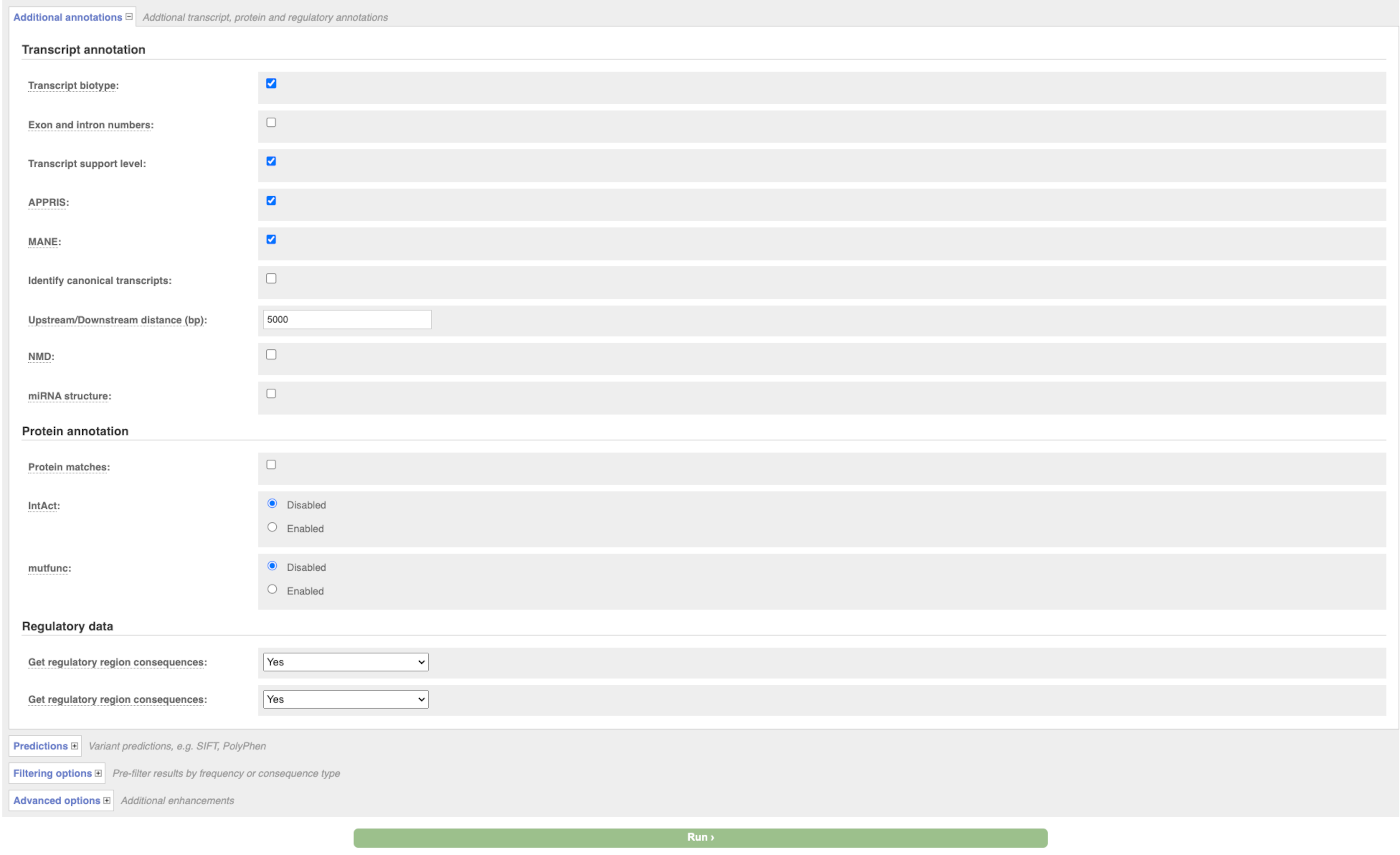
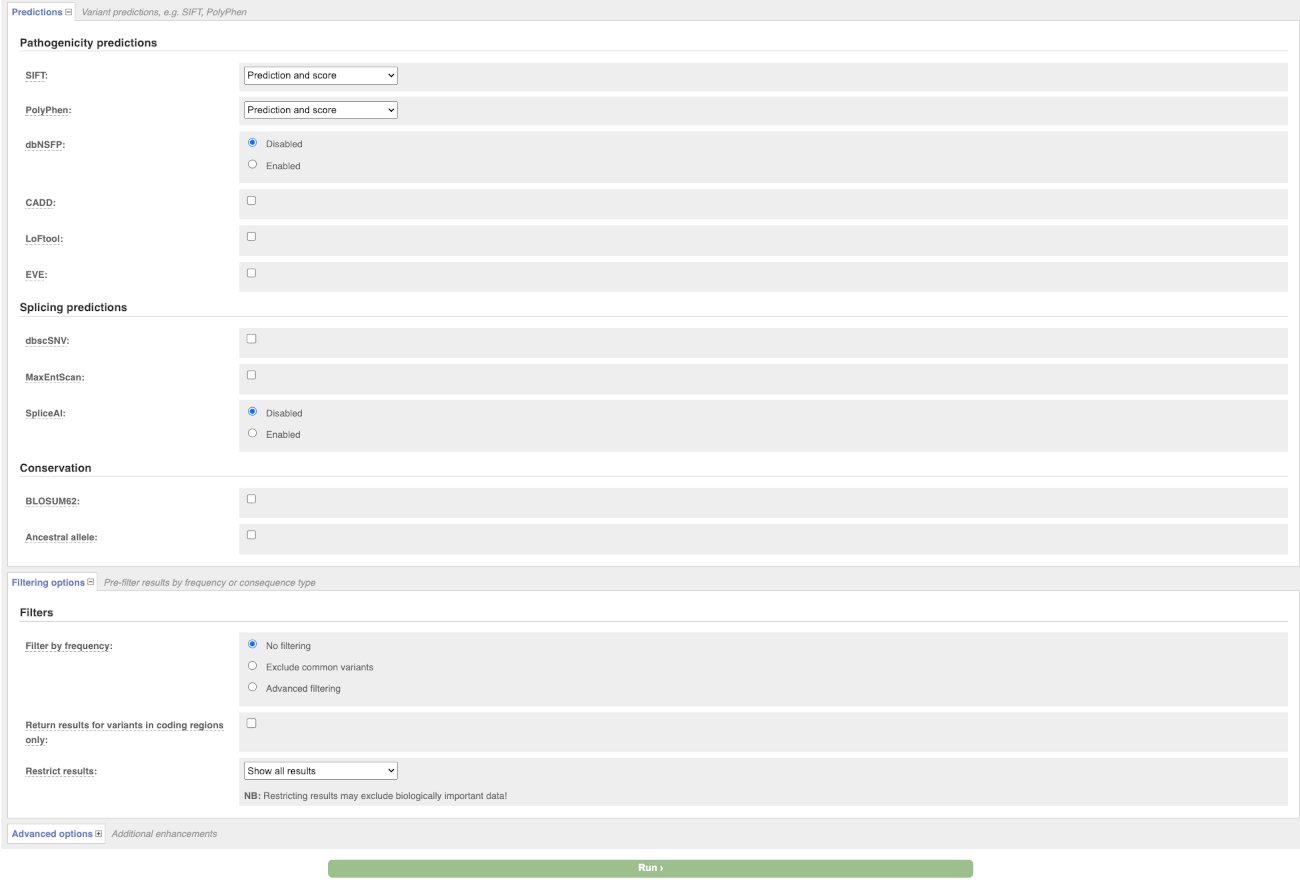
Hover over the options to see definitions.
When you have selected everything you need, scroll right to the bottom and click Run.
The display will show you the status of your job. It will say Queued, then automatically switch to Done when the job is done, you do not need to refresh the page. You can save, edit, share or delete your job at this time. If you have submitted multiple jobs, they will all appear here.

Click on View Results once your job is done.
In your results you will see a graphical and table summary of the data as well as a table with the detailed results.
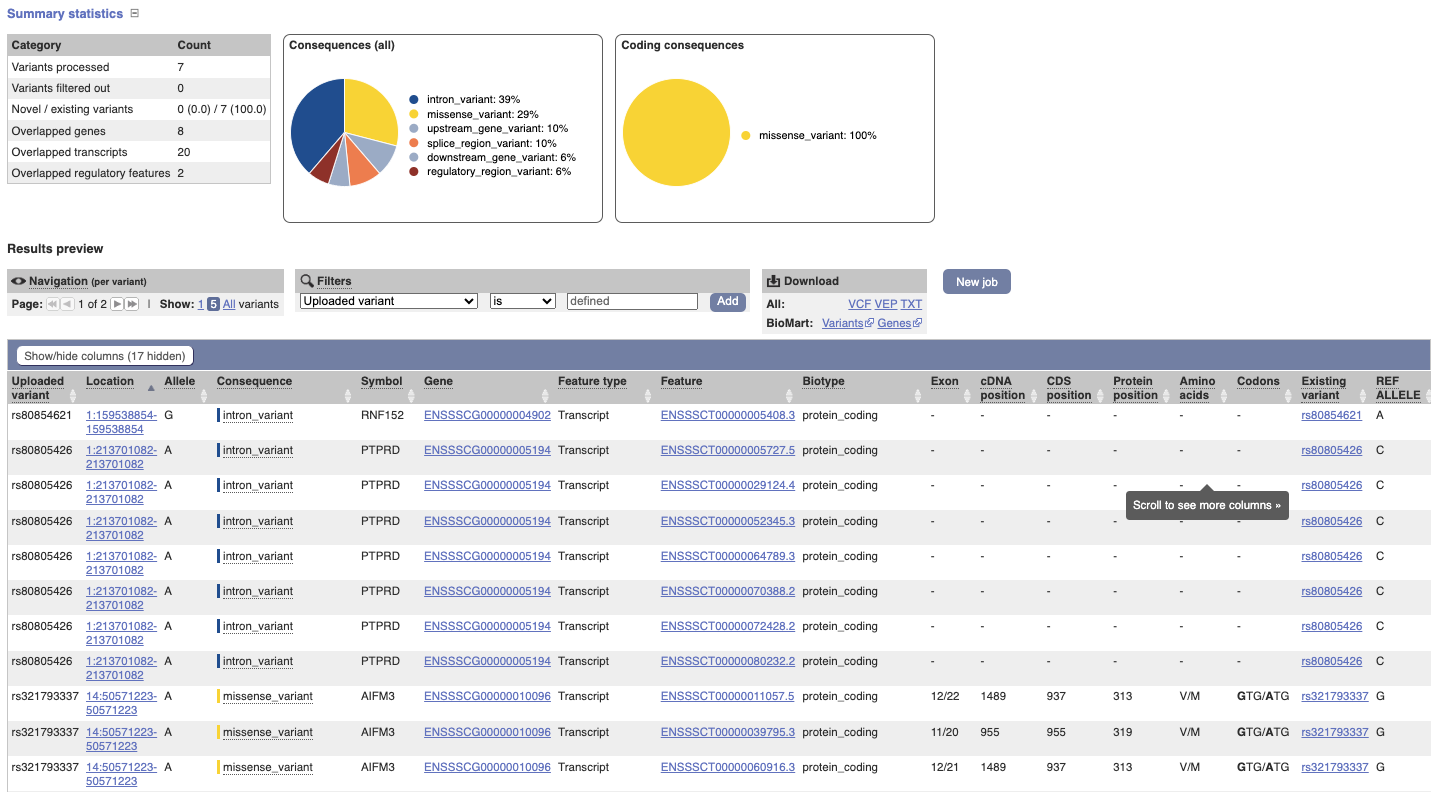
VEP for chicken data
We have identified a few variants associated with body size in chicken (bGalGal1.mat.broiler.GRCg7b):
chr 6, genomic coordinate 23650222, alleles A/C, forward strand
chr 6, genomic coordinate 23645685, alleles C/A, forward strand
chr 1, genomic coordinate 51237121, alleles C/T, forward strand
(a) Which genes and transcripts do these variants map to?
(b) What are the consequence terms for these variants?
(c) Which regulatory feature is affected by the variants?
Go to the Variant Effect Predictor (VEP) under Tools on the top banner of any Ensembl page.
Copy the following into the Paste data text box: 6 23650222 23650222 A/C + var1, 6 23645685 23645685 C/A + var2, 1 51237121 51237121 C/T + var3,
Note that this is the Ensembl default format (chr start end reference/alternate alleles). For additional formats accepted by VEP, have a look here: http://www.ensembl.org/info/docs/tools/vep/vep_formats.html
Click Run.
(a) In the Results table, you’ll see that the variants fall into three genes.
(b) The consequence terms are listed in the Consequence column and Consequences (all) chart and include intron_variant, regulatory_region_variant, upstream_gene_variant and downstream_gene_variant.
(c) Variant 3 at 1:51237121-51237121 with T allele affects regulatory feature ENSR00000006264 (promoter).







